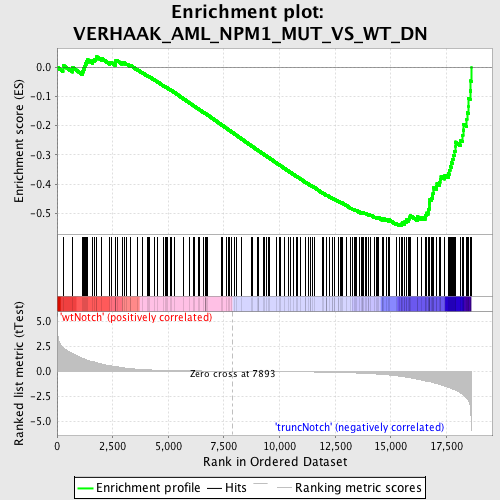
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_truncNotch_versus_wtNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch_repos |
| Phenotype | phenotype_truncNotch_versus_wtNotch.cls#wtNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | VERHAAK_AML_NPM1_MUT_VS_WT_DN |
| Enrichment Score (ES) | -0.5434037 |
| Normalized Enrichment Score (NES) | -1.6135621 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.1554414 |
| FWER p-Value | 0.958 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | POLE | 6020538 | 265 | 2.398 | 0.0067 | No | ||
| 2 | VIL2 | 5570161 | 694 | 1.755 | -0.0011 | No | ||
| 3 | ITM2A | 460576 4210086 | 1142 | 1.289 | -0.0140 | No | ||
| 4 | EMP1 | 4120438 | 1189 | 1.248 | -0.0055 | No | ||
| 5 | UGCG | 3420053 | 1230 | 1.219 | 0.0030 | No | ||
| 6 | H1F0 | 1400408 | 1259 | 1.192 | 0.0120 | No | ||
| 7 | FLNB | 2510575 | 1305 | 1.148 | 0.0196 | No | ||
| 8 | PPP1R16B | 3520300 | 1379 | 1.091 | 0.0253 | No | ||
| 9 | DNTT | 670162 | 1605 | 0.974 | 0.0216 | No | ||
| 10 | WASF1 | 2060484 | 1675 | 0.925 | 0.0260 | No | ||
| 11 | RRAGD | 670075 | 1747 | 0.879 | 0.0299 | No | ||
| 12 | GATA3 | 6130068 | 1785 | 0.855 | 0.0354 | No | ||
| 13 | RPS6KA2 | 2970161 4730278 | 2004 | 0.726 | 0.0299 | No | ||
| 14 | FOXO1A | 3120670 | 2373 | 0.553 | 0.0148 | No | ||
| 15 | MX1 | 2690301 6840364 | 2434 | 0.531 | 0.0162 | No | ||
| 16 | RAMP1 | 2320168 | 2612 | 0.467 | 0.0108 | No | ||
| 17 | ABLIM1 | 580170 3710338 6520504 | 2619 | 0.466 | 0.0145 | No | ||
| 18 | ZHX2 | 2900452 | 2635 | 0.461 | 0.0178 | No | ||
| 19 | ALDH2 | 4230019 | 2639 | 0.460 | 0.0216 | No | ||
| 20 | JUP | 2510671 | 2691 | 0.443 | 0.0228 | No | ||
| 21 | AK3L1 | 7100601 | 2922 | 0.367 | 0.0135 | No | ||
| 22 | CENTD1 | 2570592 | 2956 | 0.354 | 0.0148 | No | ||
| 23 | NEDD4 | 460736 1190280 | 3009 | 0.339 | 0.0150 | No | ||
| 24 | CD69 | 380167 4730088 | 3135 | 0.303 | 0.0109 | No | ||
| 25 | LAMC1 | 4730025 | 3306 | 0.264 | 0.0040 | No | ||
| 26 | ERG | 50154 1770739 | 3309 | 0.264 | 0.0062 | No | ||
| 27 | ARHGEF17 | 2060008 | 3632 | 0.200 | -0.0095 | No | ||
| 28 | TCF3 | 5080022 | 3817 | 0.172 | -0.0180 | No | ||
| 29 | CCR2 | 1450519 | 4059 | 0.140 | -0.0298 | No | ||
| 30 | MLLT3 | 510204 3850722 6200088 | 4097 | 0.136 | -0.0307 | No | ||
| 31 | SH3BP4 | 3140242 | 4155 | 0.131 | -0.0326 | No | ||
| 32 | GATM | 380671 | 4371 | 0.112 | -0.0433 | No | ||
| 33 | ITGA6 | 3830129 | 4521 | 0.100 | -0.0505 | No | ||
| 34 | SPARC | 1690086 | 4765 | 0.085 | -0.0629 | No | ||
| 35 | KCNE1L | 4670092 | 4856 | 0.080 | -0.0671 | No | ||
| 36 | GALC | 2260685 4810161 6370040 | 4900 | 0.077 | -0.0687 | No | ||
| 37 | HGF | 3360593 | 4957 | 0.075 | -0.0711 | No | ||
| 38 | TM6SF1 | 730181 | 5087 | 0.069 | -0.0775 | No | ||
| 39 | MPL | 2360253 | 5133 | 0.067 | -0.0794 | No | ||
| 40 | PRKD3 | 5220520 5890519 | 5270 | 0.061 | -0.0862 | No | ||
| 41 | CD48 | 1570647 | 5702 | 0.045 | -0.1092 | No | ||
| 42 | PDE4B | 4480121 | 5928 | 0.039 | -0.1211 | No | ||
| 43 | GNG7 | 1940019 | 6130 | 0.033 | -0.1317 | No | ||
| 44 | DHRS3 | 360609 | 6168 | 0.033 | -0.1334 | No | ||
| 45 | GNAI1 | 4560390 | 6354 | 0.028 | -0.1432 | No | ||
| 46 | FZD6 | 360288 | 6399 | 0.027 | -0.1453 | No | ||
| 47 | BPGM | 5080520 | 6567 | 0.024 | -0.1542 | No | ||
| 48 | DCHS1 | 4850487 | 6656 | 0.022 | -0.1588 | No | ||
| 49 | NPR3 | 1580239 | 6666 | 0.021 | -0.1591 | No | ||
| 50 | FCER1A | 1050671 | 6687 | 0.021 | -0.1600 | No | ||
| 51 | PDE3B | 2030563 | 6690 | 0.021 | -0.1599 | No | ||
| 52 | NBL1 | 2480021 | 6726 | 0.020 | -0.1616 | No | ||
| 53 | PAM | 5290528 | 6751 | 0.020 | -0.1627 | No | ||
| 54 | DACH1 | 2450593 5700592 | 7395 | 0.008 | -0.1975 | No | ||
| 55 | MST1 | 1400403 | 7424 | 0.008 | -0.1990 | No | ||
| 56 | NCAM1 | 3140026 | 7611 | 0.005 | -0.2090 | No | ||
| 57 | STK32B | 7000204 | 7701 | 0.003 | -0.2139 | No | ||
| 58 | CD34 | 6650270 | 7744 | 0.002 | -0.2161 | No | ||
| 59 | GPSM2 | 2260053 | 7849 | 0.001 | -0.2217 | No | ||
| 60 | MYH11 | 7100273 | 7968 | -0.001 | -0.2281 | No | ||
| 61 | GPA33 | 2650044 | 8073 | -0.003 | -0.2337 | No | ||
| 62 | CD38 | 6100056 | 8290 | -0.006 | -0.2454 | No | ||
| 63 | CCND2 | 5340167 | 8718 | -0.014 | -0.2685 | No | ||
| 64 | F2RL1 | 3290279 | 8792 | -0.015 | -0.2723 | No | ||
| 65 | ARHGEF12 | 3990195 | 9001 | -0.018 | -0.2834 | No | ||
| 66 | NT5E | 2030603 1400731 | 9044 | -0.019 | -0.2855 | No | ||
| 67 | OPTN | 3940041 | 9046 | -0.019 | -0.2854 | No | ||
| 68 | PTGER4 | 6400670 | 9297 | -0.023 | -0.2988 | No | ||
| 69 | AGTPBP1 | 1990735 | 9313 | -0.024 | -0.2994 | No | ||
| 70 | TRPM4 | 6980270 | 9396 | -0.025 | -0.3036 | No | ||
| 71 | GPM6B | 6220458 | 9490 | -0.027 | -0.3084 | No | ||
| 72 | EPS8 | 7050204 | 9559 | -0.028 | -0.3119 | No | ||
| 73 | PMAIP1 | 1400717 5360047 5900039 | 9849 | -0.033 | -0.3273 | No | ||
| 74 | GSN | 3830168 | 9994 | -0.036 | -0.3348 | No | ||
| 75 | BANK1 | 50750 | 10018 | -0.037 | -0.3357 | No | ||
| 76 | ST18 | 5720537 | 10237 | -0.041 | -0.3471 | No | ||
| 77 | ELA2 | 1570452 5270129 | 10402 | -0.045 | -0.3557 | No | ||
| 78 | CD19 | 6940692 | 10491 | -0.046 | -0.3600 | No | ||
| 79 | CAV1 | 870025 | 10632 | -0.050 | -0.3672 | No | ||
| 80 | CSF3R | 670500 2640239 3440091 5690020 | 10774 | -0.053 | -0.3744 | No | ||
| 81 | ROBO1 | 3710398 | 10817 | -0.054 | -0.3762 | No | ||
| 82 | ADCY3 | 2350347 3440242 | 10956 | -0.058 | -0.3831 | No | ||
| 83 | PPP4R2 | 5890035 | 11178 | -0.063 | -0.3946 | No | ||
| 84 | PTRF | 2260204 | 11182 | -0.063 | -0.3942 | No | ||
| 85 | MEF2C | 670025 780338 | 11302 | -0.067 | -0.4001 | No | ||
| 86 | IFI16 | 6520601 | 11383 | -0.069 | -0.4038 | No | ||
| 87 | MEST | 4780440 6620292 | 11405 | -0.070 | -0.4043 | No | ||
| 88 | P2RX5 | 5570427 6130435 | 11473 | -0.072 | -0.4073 | No | ||
| 89 | SPON1 | 1660301 | 11580 | -0.075 | -0.4124 | No | ||
| 90 | TKTL1 | 2810672 6760102 | 11943 | -0.088 | -0.4313 | No | ||
| 91 | RHOBTB3 | 4010687 | 11962 | -0.089 | -0.4315 | No | ||
| 92 | CLEC5A | 6420139 | 12100 | -0.095 | -0.4381 | No | ||
| 93 | TMOD1 | 3850100 | 12119 | -0.095 | -0.4382 | No | ||
| 94 | P2RY5 | 3800136 | 12129 | -0.096 | -0.4379 | No | ||
| 95 | LMO4 | 3800746 | 12260 | -0.101 | -0.4440 | No | ||
| 96 | POU4F1 | 2260315 | 12389 | -0.108 | -0.4500 | No | ||
| 97 | EHD3 | 5360484 | 12395 | -0.108 | -0.4493 | No | ||
| 98 | IL5RA | 4540091 | 12469 | -0.111 | -0.4523 | No | ||
| 99 | DPYSL2 | 2100427 3130112 5700324 | 12475 | -0.112 | -0.4516 | No | ||
| 100 | RUNX3 | 3800546 | 12636 | -0.121 | -0.4592 | No | ||
| 101 | RETN | 1050041 | 12650 | -0.121 | -0.4589 | No | ||
| 102 | CLDN15 | 4150270 4730592 | 12755 | -0.127 | -0.4634 | No | ||
| 103 | TSPAN13 | 1690450 | 12784 | -0.129 | -0.4638 | No | ||
| 104 | PALM | 130601 3850017 | 12818 | -0.131 | -0.4644 | No | ||
| 105 | FHL1 | 60278 | 12986 | -0.141 | -0.4722 | No | ||
| 106 | FGFR1 | 2190095 4570161 | 13199 | -0.156 | -0.4824 | No | ||
| 107 | TGFB1I1 | 2060288 6550450 | 13285 | -0.163 | -0.4856 | No | ||
| 108 | SLC38A1 | 520465 | 13360 | -0.168 | -0.4881 | No | ||
| 109 | TRH | 5050576 | 13408 | -0.173 | -0.4891 | No | ||
| 110 | PTGIR | 2100563 | 13461 | -0.177 | -0.4904 | No | ||
| 111 | RGS1 | 4060347 4540181 | 13580 | -0.188 | -0.4951 | No | ||
| 112 | VLDLR | 870722 3060047 5340452 6550131 | 13673 | -0.197 | -0.4984 | No | ||
| 113 | VPREB1 | 3520280 | 13725 | -0.202 | -0.4994 | No | ||
| 114 | CYP2E1 | 4480209 | 13728 | -0.203 | -0.4977 | No | ||
| 115 | PTPRM | 6370136 | 13776 | -0.208 | -0.4984 | No | ||
| 116 | SYNGR1 | 5130452 6650056 | 13845 | -0.214 | -0.5002 | No | ||
| 117 | TFPI | 1400288 5550008 | 13886 | -0.218 | -0.5005 | No | ||
| 118 | EVI1 | 2000193 2680494 | 13994 | -0.231 | -0.5043 | No | ||
| 119 | MS4A3 | 2480309 2690086 | 14006 | -0.233 | -0.5028 | No | ||
| 120 | B4GALT6 | 1850176 2340687 | 14106 | -0.243 | -0.5061 | No | ||
| 121 | MEF2A | 2350041 3360128 6040180 | 14275 | -0.266 | -0.5128 | No | ||
| 122 | APP | 2510053 | 14374 | -0.278 | -0.5157 | No | ||
| 123 | TRAF5 | 3290064 | 14379 | -0.278 | -0.5135 | No | ||
| 124 | GJA1 | 5220731 | 14454 | -0.290 | -0.5149 | No | ||
| 125 | DEPDC6 | 730326 5820239 | 14616 | -0.321 | -0.5209 | No | ||
| 126 | TSPAN7 | 870133 | 14621 | -0.322 | -0.5182 | No | ||
| 127 | PROM1 | 3170537 | 14670 | -0.329 | -0.5180 | No | ||
| 128 | SPRY1 | 450253 460093 | 14805 | -0.351 | -0.5221 | No | ||
| 129 | FGF13 | 630575 1570440 5360121 | 14902 | -0.371 | -0.5241 | No | ||
| 130 | SORL1 | 630204 | 14932 | -0.377 | -0.5223 | No | ||
| 131 | KIT | 7040095 | 15233 | -0.442 | -0.5347 | No | ||
| 132 | MMP9 | 580338 | 15380 | -0.483 | -0.5384 | No | ||
| 133 | BAALC | 5310133 6450170 | 15473 | -0.512 | -0.5389 | Yes | ||
| 134 | ICAM4 | 6770053 | 15499 | -0.521 | -0.5357 | Yes | ||
| 135 | TACSTD2 | 6660341 | 15518 | -0.529 | -0.5320 | Yes | ||
| 136 | CIITA | 2650242 | 15626 | -0.568 | -0.5328 | Yes | ||
| 137 | TNFRSF21 | 6380100 | 15633 | -0.570 | -0.5281 | Yes | ||
| 138 | SERPINE2 | 2810390 | 15692 | -0.591 | -0.5261 | Yes | ||
| 139 | HPGD | 6770192 | 15714 | -0.599 | -0.5219 | Yes | ||
| 140 | PLXND1 | 5270682 | 15807 | -0.631 | -0.5214 | Yes | ||
| 141 | CD7 | 3140465 | 15820 | -0.633 | -0.5165 | Yes | ||
| 142 | ADIPOR1 | 3140440 | 15854 | -0.644 | -0.5126 | Yes | ||
| 143 | MSLN | 2640048 | 15868 | -0.651 | -0.5076 | Yes | ||
| 144 | SCRN1 | 6040025 6580019 | 16188 | -0.791 | -0.5179 | Yes | ||
| 145 | BZRAP1 | 5290487 | 16194 | -0.794 | -0.5112 | Yes | ||
| 146 | DLK1 | 1240138 | 16382 | -0.888 | -0.5136 | Yes | ||
| 147 | FZD2 | 6480154 | 16540 | -0.962 | -0.5136 | Yes | ||
| 148 | SNCA | 5340673 | 16555 | -0.967 | -0.5059 | Yes | ||
| 149 | IGFBP7 | 520411 3060110 5290152 | 16588 | -0.984 | -0.4990 | Yes | ||
| 150 | BLNK | 70288 6590092 | 16674 | -1.009 | -0.4947 | Yes | ||
| 151 | IFITM1 | 1690601 3120438 6510075 | 16701 | -1.022 | -0.4871 | Yes | ||
| 152 | EGFL7 | 150113 670170 3840707 | 16721 | -1.029 | -0.4791 | Yes | ||
| 153 | HSPG2 | 2510687 6220750 | 16727 | -1.032 | -0.4703 | Yes | ||
| 154 | PRTN3 | 6900670 | 16734 | -1.035 | -0.4615 | Yes | ||
| 155 | MPO | 2360176 2760440 5690176 | 16754 | -1.045 | -0.4534 | Yes | ||
| 156 | CACNA2D2 | 520736 | 16839 | -1.086 | -0.4484 | Yes | ||
| 157 | PTPRCAP | 6650075 | 16872 | -1.116 | -0.4403 | Yes | ||
| 158 | ISG20 | 1770750 | 16892 | -1.129 | -0.4314 | Yes | ||
| 159 | YPEL1 | 6590435 | 16926 | -1.150 | -0.4231 | Yes | ||
| 160 | SSBP2 | 2260725 2900400 4730711 | 16927 | -1.151 | -0.4130 | Yes | ||
| 161 | LCN2 | 2510112 | 17048 | -1.231 | -0.4087 | Yes | ||
| 162 | CD3D | 2810739 | 17073 | -1.243 | -0.3990 | Yes | ||
| 163 | TRIB2 | 4120605 | 17181 | -1.321 | -0.3932 | Yes | ||
| 164 | NARFL | 2260372 4050341 | 17212 | -1.337 | -0.3831 | Yes | ||
| 165 | CHI3L1 | 1170039 | 17254 | -1.360 | -0.3734 | Yes | ||
| 166 | ITM2C | 780647 | 17428 | -1.477 | -0.3698 | Yes | ||
| 167 | ALAS2 | 6550176 | 17594 | -1.598 | -0.3647 | Yes | ||
| 168 | SERPINF1 | 7040367 | 17622 | -1.624 | -0.3519 | Yes | ||
| 169 | ASS1 | 1580632 4070082 | 17662 | -1.661 | -0.3394 | Yes | ||
| 170 | RBPMS | 3990494 | 17716 | -1.713 | -0.3272 | Yes | ||
| 171 | SERPING1 | 5550440 | 17774 | -1.761 | -0.3148 | Yes | ||
| 172 | PRODH | 1240452 | 17805 | -1.804 | -0.3006 | Yes | ||
| 173 | HSPB1 | 6760435 | 17866 | -1.859 | -0.2875 | Yes | ||
| 174 | CD2 | 430672 | 17893 | -1.885 | -0.2723 | Yes | ||
| 175 | HIST1H2BG | 5670632 | 17899 | -1.890 | -0.2560 | Yes | ||
| 176 | GMPR | 3990022 | 18136 | -2.160 | -0.2498 | Yes | ||
| 177 | ATP9A | 4780673 5390280 | 18231 | -2.297 | -0.2347 | Yes | ||
| 178 | LHFPL2 | 450020 | 18245 | -2.327 | -0.2149 | Yes | ||
| 179 | CD200 | 940725 5260647 | 18269 | -2.371 | -0.1953 | Yes | ||
| 180 | EVL | 1740113 | 18415 | -2.713 | -0.1793 | Yes | ||
| 181 | TGFBI | 2060446 6900112 | 18425 | -2.742 | -0.1557 | Yes | ||
| 182 | ZAP70 | 1410494 2260504 | 18495 | -2.950 | -0.1335 | Yes | ||
| 183 | CST7 | 380369 | 18501 | -2.985 | -0.1076 | Yes | ||
| 184 | TOX | 3440053 | 18566 | -3.421 | -0.0809 | Yes | ||
| 185 | GYPC | 2320102 | 18596 | -4.042 | -0.0470 | Yes | ||
| 186 | SIDT1 | 7100010 | 18614 | -5.462 | 0.0001 | Yes |

