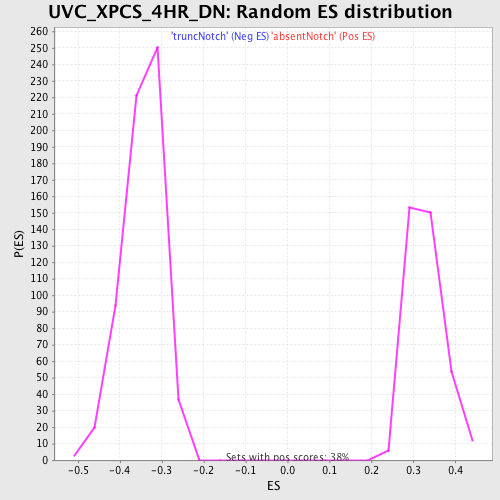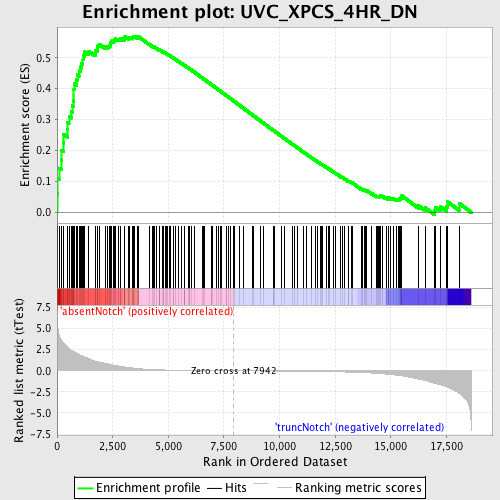
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch_repos |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#absentNotch_versus_truncNotch_repos |
| Upregulated in class | absentNotch |
| GeneSet | UVC_XPCS_4HR_DN |
| Enrichment Score (ES) | 0.5699517 |
| Normalized Enrichment Score (NES) | 1.7252246 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0402526 |
| FWER p-Value | 0.461 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PTK2 | 1780148 | 1 | 6.937 | 0.0601 | Yes | ||
| 2 | APPBP2 | 5130215 | 13 | 5.776 | 0.1096 | Yes | ||
| 3 | SRPK2 | 6380341 | 87 | 4.282 | 0.1427 | Yes | ||
| 4 | TLK1 | 7100605 | 183 | 3.696 | 0.1696 | Yes | ||
| 5 | KLF7 | 50390 | 189 | 3.660 | 0.2011 | Yes | ||
| 6 | MTAP | 2120136 | 277 | 3.321 | 0.2251 | Yes | ||
| 7 | FCHSD2 | 5720092 | 293 | 3.256 | 0.2526 | Yes | ||
| 8 | PARG | 50139 | 458 | 2.797 | 0.2679 | Yes | ||
| 9 | CUTL1 | 1570113 1770091 1850273 2100286 5340605 | 469 | 2.771 | 0.2914 | Yes | ||
| 10 | CREBBP | 5690035 7040050 | 535 | 2.625 | 0.3106 | Yes | ||
| 11 | NCOA1 | 3610438 | 637 | 2.418 | 0.3261 | Yes | ||
| 12 | RBM39 | 4230458 6450280 | 675 | 2.355 | 0.3445 | Yes | ||
| 13 | ZNF292 | 840736 1500390 | 734 | 2.263 | 0.3610 | Yes | ||
| 14 | ORC5L | 1940133 1940711 | 754 | 2.234 | 0.3793 | Yes | ||
| 15 | UGCG | 3420053 | 755 | 2.234 | 0.3987 | Yes | ||
| 16 | RASA1 | 1240315 | 794 | 2.176 | 0.4155 | Yes | ||
| 17 | EIF2C2 | 770451 | 880 | 2.077 | 0.4289 | Yes | ||
| 18 | TLK2 | 1170161 | 908 | 2.029 | 0.4450 | Yes | ||
| 19 | UPF2 | 4810471 | 985 | 1.906 | 0.4575 | Yes | ||
| 20 | RUNX1 | 3840711 | 1043 | 1.837 | 0.4703 | Yes | ||
| 21 | TLE1 | 540390 1580309 3710019 | 1108 | 1.770 | 0.4822 | Yes | ||
| 22 | APRIN | 2810050 | 1162 | 1.712 | 0.4941 | Yes | ||
| 23 | CHD1 | 4590048 | 1176 | 1.692 | 0.5081 | Yes | ||
| 24 | CENTG2 | 4210053 | 1236 | 1.626 | 0.5190 | Yes | ||
| 25 | TIAM1 | 5420288 | 1420 | 1.424 | 0.5214 | Yes | ||
| 26 | MARK3 | 2370450 | 1711 | 1.145 | 0.5156 | Yes | ||
| 27 | ITSN2 | 780484 | 1720 | 1.139 | 0.5251 | Yes | ||
| 28 | MECP2 | 1940450 | 1803 | 1.075 | 0.5300 | Yes | ||
| 29 | REV3L | 1090717 | 1823 | 1.065 | 0.5382 | Yes | ||
| 30 | SEC24B | 460717 3060040 | 1885 | 1.021 | 0.5437 | Yes | ||
| 31 | BHLHB2 | 7040603 | 2152 | 0.878 | 0.5369 | Yes | ||
| 32 | NCOA3 | 4540195 | 2284 | 0.802 | 0.5367 | Yes | ||
| 33 | DYRK1A | 3190181 | 2353 | 0.763 | 0.5397 | Yes | ||
| 34 | RNGTT | 780152 780746 1090471 | 2390 | 0.749 | 0.5442 | Yes | ||
| 35 | PPP1R12A | 4070066 4590162 | 2393 | 0.748 | 0.5506 | Yes | ||
| 36 | CNOT4 | 2320242 6760706 | 2440 | 0.713 | 0.5543 | Yes | ||
| 37 | TRIM33 | 580619 2230280 3990433 6200747 | 2532 | 0.659 | 0.5551 | Yes | ||
| 38 | SOS2 | 2060722 | 2589 | 0.639 | 0.5576 | Yes | ||
| 39 | SMURF2 | 1940161 | 2624 | 0.622 | 0.5611 | Yes | ||
| 40 | ANKRD17 | 2120402 3850035 6400044 | 2741 | 0.565 | 0.5597 | Yes | ||
| 41 | THRAP2 | 4920600 | 2843 | 0.524 | 0.5588 | Yes | ||
| 42 | IFRD1 | 4590215 | 2856 | 0.519 | 0.5626 | Yes | ||
| 43 | ZNF148 | 6420403 | 3012 | 0.459 | 0.5582 | Yes | ||
| 44 | STRN3 | 1450093 6200685 | 3021 | 0.454 | 0.5617 | Yes | ||
| 45 | ARFGEF1 | 6760494 | 3031 | 0.452 | 0.5651 | Yes | ||
| 46 | CTBP2 | 430309 3710079 | 3047 | 0.445 | 0.5682 | Yes | ||
| 47 | MARCH7 | 3800020 6620168 | 3224 | 0.383 | 0.5620 | Yes | ||
| 48 | ATF2 | 2260348 2480441 | 3226 | 0.383 | 0.5652 | Yes | ||
| 49 | NRIP1 | 3190039 | 3269 | 0.369 | 0.5662 | Yes | ||
| 50 | MYC | 380541 4670170 | 3372 | 0.337 | 0.5636 | Yes | ||
| 51 | BMPR1A | 2190193 4060603 | 3376 | 0.335 | 0.5663 | Yes | ||
| 52 | CREB1 | 1500717 2230358 3610600 6550601 | 3414 | 0.323 | 0.5671 | Yes | ||
| 53 | MALT1 | 4670292 | 3450 | 0.311 | 0.5679 | Yes | ||
| 54 | STK3 | 7100427 | 3462 | 0.306 | 0.5700 | Yes | ||
| 55 | BARD1 | 3170348 | 3612 | 0.261 | 0.5641 | No | ||
| 56 | FNDC3A | 2690097 3290044 | 3635 | 0.255 | 0.5652 | No | ||
| 57 | ZFYVE9 | 6980288 7100017 | 3637 | 0.254 | 0.5673 | No | ||
| 58 | TCF7L2 | 2190348 5220113 | 3667 | 0.248 | 0.5679 | No | ||
| 59 | MYO10 | 3830576 5570092 | 4154 | 0.153 | 0.5429 | No | ||
| 60 | PIP5K3 | 5360112 | 4307 | 0.134 | 0.5358 | No | ||
| 61 | THRAP1 | 2470121 | 4332 | 0.131 | 0.5356 | No | ||
| 62 | STARD13 | 540722 | 4364 | 0.128 | 0.5351 | No | ||
| 63 | ACVR1 | 6840671 | 4475 | 0.115 | 0.5301 | No | ||
| 64 | PTPRK | 1450433 2640142 | 4579 | 0.106 | 0.5254 | No | ||
| 65 | TOPBP1 | 6020333 | 4595 | 0.105 | 0.5255 | No | ||
| 66 | STAM | 3850008 4760441 | 4604 | 0.104 | 0.5260 | No | ||
| 67 | DAPK1 | 5570010 | 4744 | 0.093 | 0.5193 | No | ||
| 68 | KLF6 | 110541 5360026 | 4763 | 0.092 | 0.5191 | No | ||
| 69 | FYN | 2100468 4760520 4850687 | 4785 | 0.091 | 0.5187 | No | ||
| 70 | PTPN12 | 2030309 6020725 6130746 6290609 | 4851 | 0.085 | 0.5160 | No | ||
| 71 | RCOR1 | 5360717 | 4910 | 0.082 | 0.5135 | No | ||
| 72 | IL6 | 380133 | 4953 | 0.080 | 0.5119 | No | ||
| 73 | BLM | 520619 5570170 | 5035 | 0.075 | 0.5082 | No | ||
| 74 | ACSL3 | 3140195 | 5101 | 0.072 | 0.5053 | No | ||
| 75 | IRAK1BP1 | 610037 | 5211 | 0.066 | 0.5000 | No | ||
| 76 | GNAI1 | 4560390 | 5227 | 0.065 | 0.4997 | No | ||
| 77 | MYST4 | 1400563 2570687 3360458 6840402 | 5336 | 0.061 | 0.4944 | No | ||
| 78 | F3 | 2940180 | 5476 | 0.056 | 0.4873 | No | ||
| 79 | BICD1 | 110128 2940324 | 5568 | 0.053 | 0.4829 | No | ||
| 80 | AKAP9 | 510184 2060195 2470136 4280048 | 5709 | 0.048 | 0.4757 | No | ||
| 81 | IGF1R | 3360494 | 5719 | 0.048 | 0.4756 | No | ||
| 82 | ZCCHC11 | 3390731 5360039 | 5726 | 0.048 | 0.4757 | No | ||
| 83 | TGFBR3 | 5290577 | 5923 | 0.042 | 0.4654 | No | ||
| 84 | TSPAN5 | 6770019 | 5950 | 0.041 | 0.4644 | No | ||
| 85 | FGF5 | 5270204 | 5954 | 0.041 | 0.4646 | No | ||
| 86 | ARHGAP12 | 1050181 | 6060 | 0.038 | 0.4592 | No | ||
| 87 | TGFB2 | 4920292 | 6193 | 0.034 | 0.4524 | No | ||
| 88 | SCHIP1 | 4560152 | 6555 | 0.026 | 0.4330 | No | ||
| 89 | TIPARP | 50397 | 6561 | 0.026 | 0.4330 | No | ||
| 90 | EHBP1 | 430348 | 6630 | 0.024 | 0.4295 | No | ||
| 91 | VEGFC | 5910494 | 6922 | 0.018 | 0.4139 | No | ||
| 92 | CLASP2 | 2510139 | 6949 | 0.018 | 0.4126 | No | ||
| 93 | GLRB | 2510433 4280736 | 6999 | 0.017 | 0.4101 | No | ||
| 94 | E2F5 | 5860575 | 7163 | 0.014 | 0.4014 | No | ||
| 95 | USP15 | 610592 3520504 | 7236 | 0.012 | 0.3976 | No | ||
| 96 | TLE4 | 1450064 1500519 | 7323 | 0.010 | 0.3930 | No | ||
| 97 | ARHGEF7 | 1690441 | 7397 | 0.009 | 0.3891 | No | ||
| 98 | PVRL3 | 1400324 4200593 6650128 | 7610 | 0.005 | 0.3777 | No | ||
| 99 | PLEKHC1 | 2650066 | 7620 | 0.005 | 0.3773 | No | ||
| 100 | PUM2 | 4200441 5910446 | 7693 | 0.004 | 0.3734 | No | ||
| 101 | PCTK2 | 5700673 | 7791 | 0.002 | 0.3681 | No | ||
| 102 | KLF10 | 4850056 | 7935 | 0.000 | 0.3604 | No | ||
| 103 | SHB | 4920494 | 7988 | -0.001 | 0.3576 | No | ||
| 104 | NUP153 | 7000452 | 8186 | -0.004 | 0.3469 | No | ||
| 105 | SAMD4A | 1170717 | 8358 | -0.007 | 0.3377 | No | ||
| 106 | SATB2 | 2470181 | 8760 | -0.014 | 0.3161 | No | ||
| 107 | FBXW11 | 6450632 | 8838 | -0.015 | 0.3121 | No | ||
| 108 | CTGF | 4540577 | 9137 | -0.020 | 0.2961 | No | ||
| 109 | RGS7 | 3390044 | 9297 | -0.023 | 0.2877 | No | ||
| 110 | SRGAP2 | 1050092 1230403 3830066 6130215 | 9714 | -0.031 | 0.2654 | No | ||
| 111 | TAF4 | 1450170 | 9716 | -0.031 | 0.2656 | No | ||
| 112 | WDR37 | 2630519 6840347 | 9760 | -0.032 | 0.2636 | No | ||
| 113 | TRIM2 | 430546 1170286 | 10102 | -0.039 | 0.2454 | No | ||
| 114 | RNF13 | 2370021 | 10237 | -0.042 | 0.2385 | No | ||
| 115 | CENPE | 2850022 | 10578 | -0.049 | 0.2205 | No | ||
| 116 | FBXL7 | 6620711 | 10661 | -0.051 | 0.2165 | No | ||
| 117 | TERF1 | 2340113 | 10806 | -0.055 | 0.2092 | No | ||
| 118 | CENPC1 | 610273 | 11074 | -0.061 | 0.1952 | No | ||
| 119 | CNOT2 | 520546 2060397 4610543 5550113 | 11210 | -0.064 | 0.1885 | No | ||
| 120 | GNAQ | 430670 4210131 5900736 | 11427 | -0.072 | 0.1774 | No | ||
| 121 | PHLPP | 3440563 | 11629 | -0.078 | 0.1672 | No | ||
| 122 | CENTD1 | 2570592 | 11719 | -0.081 | 0.1630 | No | ||
| 123 | DDX10 | 520746 | 11849 | -0.086 | 0.1568 | No | ||
| 124 | MID1 | 4070114 4760458 | 11900 | -0.089 | 0.1549 | No | ||
| 125 | NFIB | 460450 | 11926 | -0.089 | 0.1543 | No | ||
| 126 | SOS1 | 7050338 | 12089 | -0.096 | 0.1463 | No | ||
| 127 | NCK1 | 6200575 6510050 | 12187 | -0.100 | 0.1419 | No | ||
| 128 | BCAR3 | 1660717 | 12228 | -0.102 | 0.1407 | No | ||
| 129 | EPHA4 | 460750 | 12418 | -0.113 | 0.1314 | No | ||
| 130 | KIF2A | 3990286 6130575 | 12527 | -0.119 | 0.1266 | No | ||
| 131 | DNMBP | 2810368 6900403 | 12757 | -0.134 | 0.1153 | No | ||
| 132 | CYR61 | 1240408 5290026 4120452 6550008 | 12825 | -0.138 | 0.1129 | No | ||
| 133 | TUBGCP3 | 4920563 | 12924 | -0.145 | 0.1088 | No | ||
| 134 | IL1RAP | 50546 450711 1580397 3940301 5890181 6370373 | 13116 | -0.159 | 0.0999 | No | ||
| 135 | CD2AP | 1940369 | 13118 | -0.159 | 0.1012 | No | ||
| 136 | ZZZ3 | 780133 2320239 2640452 5080706 6550341 | 13217 | -0.167 | 0.0973 | No | ||
| 137 | WNT5A | 840685 3120152 | 13276 | -0.171 | 0.0957 | No | ||
| 138 | PAFAH1B1 | 4230333 6420121 6450066 | 13670 | -0.213 | 0.0762 | No | ||
| 139 | CENPA | 5080154 | 13738 | -0.221 | 0.0745 | No | ||
| 140 | DOCK4 | 5910102 | 13805 | -0.229 | 0.0729 | No | ||
| 141 | PBX3 | 1300424 3710577 6180575 | 13878 | -0.236 | 0.0710 | No | ||
| 142 | SPEN | 2060041 | 13919 | -0.241 | 0.0710 | No | ||
| 143 | RRAS2 | 6290170 | 14142 | -0.272 | 0.0613 | No | ||
| 144 | EXT1 | 4540603 | 14352 | -0.307 | 0.0526 | No | ||
| 145 | HOXB2 | 6450592 | 14396 | -0.314 | 0.0530 | No | ||
| 146 | FGF2 | 3450433 | 14449 | -0.323 | 0.0530 | No | ||
| 147 | MICAL3 | 6040333 | 14510 | -0.333 | 0.0526 | No | ||
| 148 | SOCS5 | 3830398 7100093 | 14544 | -0.339 | 0.0538 | No | ||
| 149 | EGFR | 4920138 6480521 | 14611 | -0.351 | 0.0533 | No | ||
| 150 | CBLB | 3520465 4670348 | 14787 | -0.390 | 0.0471 | No | ||
| 151 | CDYL | 4200142 4730397 6060400 | 14892 | -0.411 | 0.0451 | No | ||
| 152 | ANKS1A | 50148 | 14906 | -0.415 | 0.0480 | No | ||
| 153 | MTF2 | 50333 | 15006 | -0.438 | 0.0464 | No | ||
| 154 | SWAP70 | 2810577 2850739 5130594 | 15100 | -0.466 | 0.0454 | No | ||
| 155 | DLC1 | 1090632 6450594 | 15250 | -0.516 | 0.0418 | No | ||
| 156 | CDK8 | 1050113 1190601 1780358 6110102 | 15335 | -0.544 | 0.0420 | No | ||
| 157 | ZHX2 | 2900452 | 15396 | -0.567 | 0.0436 | No | ||
| 158 | ZHX3 | 460615 | 15422 | -0.576 | 0.0472 | No | ||
| 159 | HIRA | 4480609 5270167 | 15491 | -0.601 | 0.0488 | No | ||
| 160 | AKAP10 | 2630504 5910577 | 15499 | -0.603 | 0.0536 | No | ||
| 161 | BTRC | 3170131 | 16231 | -0.978 | 0.0225 | No | ||
| 162 | RBM16 | 6290286 6400181 | 16548 | -1.162 | 0.0154 | No | ||
| 163 | ATXN2 | 3940017 4810242 | 16954 | -1.473 | 0.0062 | No | ||
| 164 | MTMR3 | 1050014 4920180 | 16991 | -1.493 | 0.0172 | No | ||
| 165 | MYST3 | 5270500 | 17220 | -1.648 | 0.0192 | No | ||
| 166 | LHFPL2 | 450020 | 17492 | -1.894 | 0.0209 | No | ||
| 167 | SLC25A12 | 7400731 | 17566 | -1.964 | 0.0340 | No | ||
| 168 | PRR3 | 3440112 | 18069 | -2.642 | 0.0296 | No |