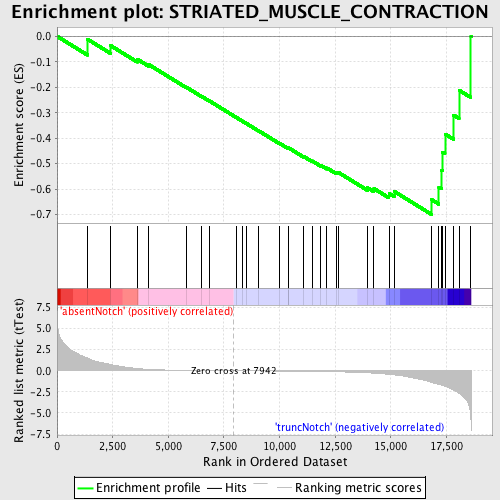
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch_repos |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#absentNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | STRIATED_MUSCLE_CONTRACTION |
| Enrichment Score (ES) | -0.699526 |
| Normalized Enrichment Score (NES) | -1.5876181 |
| Nominal p-value | 0.0037105752 |
| FDR q-value | 0.31830934 |
| FWER p-Value | 0.984 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TPM4 | 2100059 3850072 4920162 | 1367 | 1.480 | -0.0115 | No | ||
| 2 | MYH3 | 110400 | 2412 | 0.732 | -0.0369 | No | ||
| 3 | TPM2 | 520735 3870390 | 3603 | 0.264 | -0.0898 | No | ||
| 4 | ACTA2 | 60008 1230082 | 4128 | 0.157 | -0.1114 | No | ||
| 5 | MYL3 | 6040563 | 5814 | 0.045 | -0.2002 | No | ||
| 6 | MYL1 | 2450707 | 6502 | 0.027 | -0.2361 | No | ||
| 7 | MYL9 | 4210750 7050138 | 6857 | 0.019 | -0.2543 | No | ||
| 8 | MYL2 | 70471 | 8047 | -0.001 | -0.3182 | No | ||
| 9 | MYOM1 | 580091 | 8338 | -0.007 | -0.3336 | No | ||
| 10 | NEB | 580735 | 8506 | -0.009 | -0.3421 | No | ||
| 11 | MYH6 | 2900373 | 9058 | -0.019 | -0.3710 | No | ||
| 12 | MYBPC2 | 1980368 | 10013 | -0.037 | -0.4208 | No | ||
| 13 | DMD | 1740041 3990332 | 10389 | -0.045 | -0.4391 | No | ||
| 14 | DES | 1450341 | 10402 | -0.045 | -0.4378 | No | ||
| 15 | ACTA1 | 840538 | 11083 | -0.061 | -0.4719 | No | ||
| 16 | TNNT2 | 2450364 | 11468 | -0.073 | -0.4894 | No | ||
| 17 | TMOD1 | 3850100 | 11860 | -0.087 | -0.5068 | No | ||
| 18 | TTN | 2320161 4670056 6550026 | 12124 | -0.098 | -0.5169 | No | ||
| 19 | TNNT1 | 3360671 | 12560 | -0.121 | -0.5352 | No | ||
| 20 | TNNT3 | 1570239 | 12629 | -0.124 | -0.5337 | No | ||
| 21 | MYBPC3 | 650390 | 13956 | -0.246 | -0.5947 | No | ||
| 22 | TNNC2 | 3840079 | 14241 | -0.287 | -0.5979 | No | ||
| 23 | ACTN3 | 3140541 6480598 | 14918 | -0.418 | -0.6167 | No | ||
| 24 | VIM | 20431 | 15151 | -0.483 | -0.6089 | No | ||
| 25 | TNNI1 | 3710040 | 16836 | -1.380 | -0.6416 | Yes | ||
| 26 | TNNI3 | 7000093 | 17160 | -1.601 | -0.5918 | Yes | ||
| 27 | TPM1 | 130673 | 17289 | -1.716 | -0.5266 | Yes | ||
| 28 | ACTN2 | 4200435 | 17324 | -1.743 | -0.4553 | Yes | ||
| 29 | TNNI2 | 5910739 | 17436 | -1.850 | -0.3836 | Yes | ||
| 30 | TCAP | 4890446 | 17816 | -2.264 | -0.3090 | Yes | ||
| 31 | ACTN4 | 3840301 4590390 7050132 | 18067 | -2.640 | -0.2116 | Yes | ||
| 32 | MYL4 | 6860288 | 18601 | -5.744 | 0.0008 | Yes |

