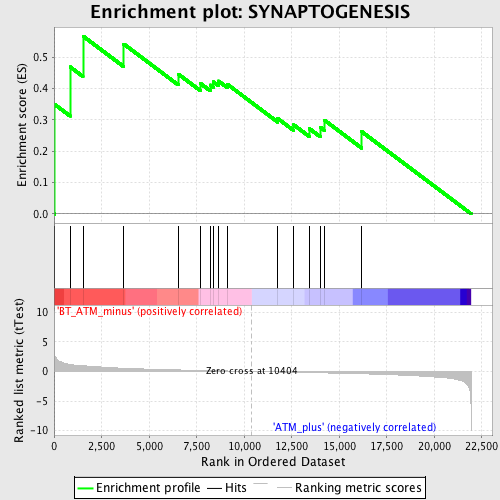
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | SYNAPTOGENESIS |
| Enrichment Score (ES) | 0.5676684 |
| Normalized Enrichment Score (NES) | 1.5728576 |
| Nominal p-value | 0.04112554 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PCDHB4 | 1422951_at 1440632_at | 43 | 2.617 | 0.3488 | Yes | ||
| 2 | POU4F1 | 1429667_at 1429668_at | 843 | 1.172 | 0.4694 | Yes | ||
| 3 | PCDHB11 | 1450270_at 1456710_at | 1521 | 0.963 | 0.5677 | Yes | ||
| 4 | PCDHB3 | 1420429_at | 3664 | 0.540 | 0.5424 | No | ||
| 5 | ACHE | 1422635_at | 6529 | 0.258 | 0.4463 | No | ||
| 6 | PCDHB10 | 1422729_at | 7701 | 0.174 | 0.4162 | No | ||
| 7 | PCDHB13 | 1450316_at | 8211 | 0.139 | 0.4116 | No | ||
| 8 | PCDHB5 | 1450263_at | 8372 | 0.128 | 0.4215 | No | ||
| 9 | PCDHB6 | 1423014_at | 8639 | 0.112 | 0.4244 | No | ||
| 10 | COL4A4 | 1425772_at 1440250_at 1445328_at | 9108 | 0.083 | 0.4141 | No | ||
| 11 | PCDHB14 | 1421437_x_at | 11764 | -0.085 | 0.3043 | No | ||
| 12 | PCDHB9 | 1422640_at | 12606 | -0.136 | 0.2842 | No | ||
| 13 | NLGN1 | 1421648_at 1437160_at | 13439 | -0.190 | 0.2717 | No | ||
| 14 | PCDHB16 | 1441358_at 1441994_at 1450216_at | 13996 | -0.227 | 0.2768 | No | ||
| 15 | GHRL | 1448980_at | 14238 | -0.244 | 0.2985 | No | ||
| 16 | PCDHB2 | 1421548_at | 16180 | -0.392 | 0.2625 | No |

