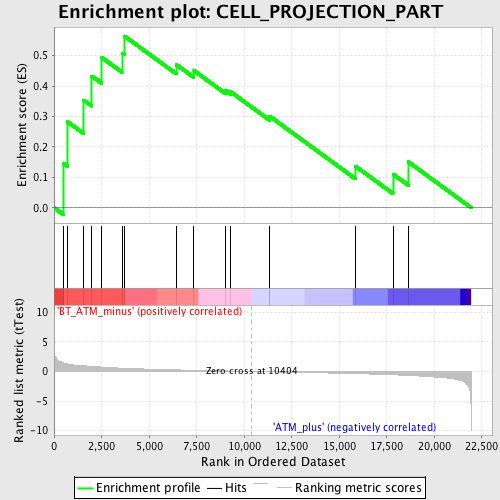
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | CELL_PROJECTION_PART |
| Enrichment Score (ES) | 0.5640528 |
| Normalized Enrichment Score (NES) | 1.549688 |
| Nominal p-value | 0.028199567 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | B4GALT1 | 1418014_a_at 1458616_at 1458773_at | 483 | 1.470 | 0.1463 | Yes | ||
| 2 | SOD1 | 1435304_at 1440222_at 1440896_at 1451124_at 1459976_s_at | 693 | 1.278 | 0.2832 | Yes | ||
| 3 | SYNPO | 1427045_at 1434089_at | 1560 | 0.957 | 0.3533 | Yes | ||
| 4 | CABP4 | 1425878_at 1456936_at | 1977 | 0.861 | 0.4329 | Yes | ||
| 5 | CDK5R1 | 1421123_at 1421124_at 1433450_at 1433451_at | 2504 | 0.741 | 0.4938 | Yes | ||
| 6 | ACTN2 | 1448327_at 1456968_at | 3572 | 0.553 | 0.5085 | Yes | ||
| 7 | PKHD1 | 1419820_at | 3700 | 0.536 | 0.5641 | Yes | ||
| 8 | SLC22A12 | 1422897_at 1422898_s_at | 6429 | 0.267 | 0.4702 | No | ||
| 9 | DNAH9 | 1442412_at 1457401_at | 7354 | 0.198 | 0.4507 | No | ||
| 10 | ATP6V0A4 | 1422030_at 1443732_at | 9018 | 0.088 | 0.3850 | No | ||
| 11 | PKD2 | 1417753_at 1441870_s_at | 9279 | 0.071 | 0.3813 | No | ||
| 12 | SPAG6 | 1417528_at | 11330 | -0.058 | 0.2944 | No | ||
| 13 | CROCC | 1427338_at | 11334 | -0.058 | 0.3009 | No | ||
| 14 | DNALI1 | 1436126_at 1455379_at | 15837 | -0.363 | 0.1371 | No | ||
| 15 | EFHC1 | 1453159_at | 17826 | -0.560 | 0.1105 | No | ||
| 16 | RAB35 | 1433922_at | 18628 | -0.671 | 0.1508 | No |

