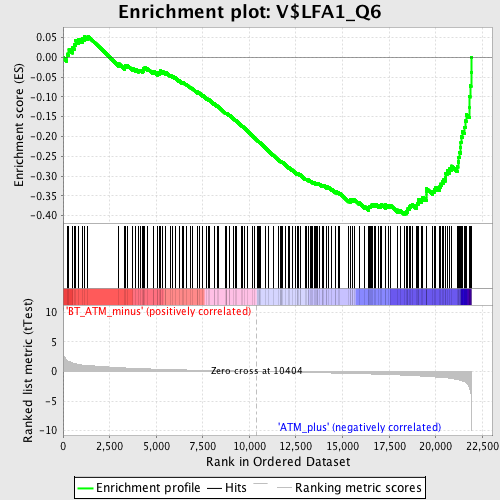
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | V$LFA1_Q6 |
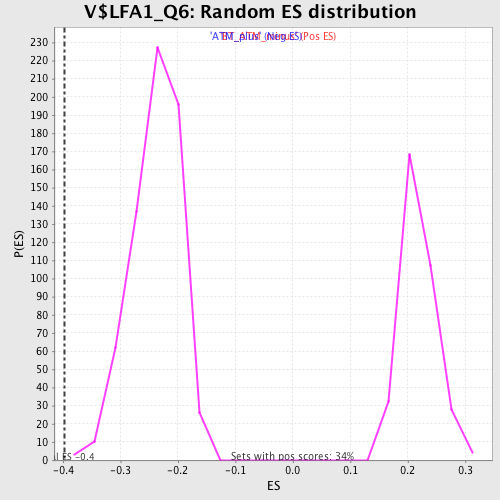
| Enrichment Score (ES) | -0.39718103 |
| Normalized Enrichment Score (NES) | -1.6640375 |
| Nominal p-value | 0.0015128592 |
| FDR q-value | 0.039609414 |
| FWER p-Value | 0.41 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SMAD3 | 1450471_at 1450472_s_at 1454960_at | 208 | 1.878 | 0.0085 | No | ||
| 2 | NR5A2 | 1420410_at 1449706_s_at 1449707_at | 315 | 1.655 | 0.0196 | No | ||
| 3 | ASXL1 | 1434561_at 1435077_at 1458380_at | 504 | 1.443 | 0.0248 | No | ||
| 4 | HES1 | 1418102_at | 594 | 1.348 | 0.0337 | No | ||
| 5 | TPCN1 | 1434930_at 1451771_at 1451772_at | 672 | 1.287 | 0.0426 | No | ||
| 6 | POU4F1 | 1429667_at 1429668_at | 843 | 1.172 | 0.0460 | No | ||
| 7 | RAB31 | 1416165_at 1431691_a_at | 1031 | 1.080 | 0.0478 | No | ||
| 8 | CLTC | 1419407_at 1439094_at 1440457_at 1454626_at | 1139 | 1.047 | 0.0530 | No | ||
| 9 | ACVR1C | 1428032_at 1443225_at | 1334 | 1.002 | 0.0537 | No | ||
| 10 | HNRPR | 1427129_a_at 1438807_at 1452030_a_at | 2997 | 0.647 | -0.0165 | No | ||
| 11 | BCL11B | 1440137_at 1441762_at 1450339_a_at | 3305 | 0.595 | -0.0249 | No | ||
| 12 | GPR3 | 1460275_at | 3344 | 0.588 | -0.0210 | No | ||
| 13 | ADM | 1416077_at 1447839_x_at | 3456 | 0.569 | -0.0206 | No | ||
| 14 | CHX10 | 1419628_at | 3726 | 0.531 | -0.0278 | No | ||
| 15 | GRIN2A | 1421616_at | 3884 | 0.512 | -0.0301 | No | ||
| 16 | NEURL | 1422251_at 1451872_a_at 1453527_a_at 1456854_at 1458561_at 1460343_at | 4040 | 0.495 | -0.0325 | No | ||
| 17 | NPTXR | 1421181_at 1434270_at 1450147_at | 4135 | 0.480 | -0.0322 | No | ||
| 18 | BAX | 1416837_at | 4273 | 0.465 | -0.0340 | No | ||
| 19 | PALM | 1422034_a_at 1423967_at 1439927_at 1457217_at | 4305 | 0.461 | -0.0310 | No | ||
| 20 | CTGF | 1416953_at | 4313 | 0.461 | -0.0269 | No | ||
| 21 | CEBPB | 1418901_at 1427844_a_at | 4394 | 0.452 | -0.0262 | No | ||
| 22 | RBPSUH | 1418114_at 1438848_at 1446436_at 1448957_at 1454896_at | 4547 | 0.435 | -0.0290 | No | ||
| 23 | ZNF385 | 1418865_at | 4830 | 0.407 | -0.0381 | No | ||
| 24 | PDLIM7 | 1417959_at 1428319_at 1443785_x_at | 4843 | 0.405 | -0.0347 | No | ||
| 25 | UBL3 | 1423461_a_at | 5068 | 0.384 | -0.0413 | No | ||
| 26 | PNOC | 1425892_a_at | 5071 | 0.384 | -0.0377 | No | ||
| 27 | CCNL1 | 1423622_a_at 1437383_at | 5181 | 0.373 | -0.0391 | No | ||
| 28 | SLC13A5 | 1459729_at | 5207 | 0.371 | -0.0367 | No | ||
| 29 | LZTS2 | 1426409_at | 5213 | 0.370 | -0.0334 | No | ||
| 30 | EPHB2 | 1425015_at 1425016_at 1442471_at 1445417_at 1454022_at | 5355 | 0.357 | -0.0364 | No | ||
| 31 | FLT1 | 1419300_at 1440926_at 1451756_at 1454037_a_at | 5503 | 0.344 | -0.0399 | No | ||
| 32 | KCNQ4 | 1435721_at 1442909_at | 5523 | 0.343 | -0.0374 | No | ||
| 33 | M6PR | 1416385_a_at 1416386_a_at | 5767 | 0.321 | -0.0455 | No | ||
| 34 | NYX | 1446344_at | 5895 | 0.308 | -0.0484 | No | ||
| 35 | GAD1 | 1416561_at 1416562_at | 6026 | 0.298 | -0.0515 | No | ||
| 36 | SP7 | 1418425_at | 6247 | 0.280 | -0.0589 | No | ||
| 37 | NEUROG1 | 1438551_at 1450836_at | 6427 | 0.267 | -0.0646 | No | ||
| 38 | NPAS3 | 1421567_at 1429138_at 1438060_at 1450287_at | 6462 | 0.264 | -0.0636 | No | ||
| 39 | RAB5C | 1424684_at | 6605 | 0.252 | -0.0677 | No | ||
| 40 | PPP1R16A | 1452724_at 1459751_s_at | 6819 | 0.237 | -0.0752 | No | ||
| 41 | TBR1 | 1416711_at | 6927 | 0.228 | -0.0779 | No | ||
| 42 | SIX4 | 1425767_a_at 1427701_a_at 1439753_x_at 1456862_at | 7205 | 0.208 | -0.0887 | No | ||
| 43 | PDYN | 1416266_at | 7214 | 0.207 | -0.0871 | No | ||
| 44 | GSS | 1441931_x_at 1448273_at | 7320 | 0.201 | -0.0900 | No | ||
| 45 | TNRC4 | 1452851_at | 7499 | 0.189 | -0.0963 | No | ||
| 46 | CAV1 | 1449145_a_at | 7723 | 0.172 | -0.1049 | No | ||
| 47 | OLFML2A | 1441635_at 1455743_at | 7830 | 0.166 | -0.1082 | No | ||
| 48 | ACCN1 | 1417994_a_at | 7850 | 0.164 | -0.1075 | No | ||
| 49 | KIF1C | 1424747_at 1426325_at | 8111 | 0.146 | -0.1180 | No | ||
| 50 | S100A5 | 1449927_at | 8127 | 0.145 | -0.1173 | No | ||
| 51 | HOXC6 | 1427361_at 1427362_x_at 1427454_at | 8275 | 0.134 | -0.1228 | No | ||
| 52 | PLA2G10 | 1451502_at | 8343 | 0.130 | -0.1246 | No | ||
| 53 | UQCRC1 | 1428782_a_at | 8734 | 0.105 | -0.1415 | No | ||
| 54 | HBLD1 | 1424154_a_at | 8760 | 0.104 | -0.1417 | No | ||
| 55 | NFAT5 | 1426029_a_at 1438999_a_at 1439328_at 1439805_at 1448966_a_at 1451921_a_at 1458213_at | 8761 | 0.104 | -0.1407 | No | ||
| 56 | FES | 1427368_x_at 1452410_a_at | 8787 | 0.102 | -0.1409 | No | ||
| 57 | PRRX1 | 1425526_a_at 1425527_at 1425528_at 1432129_a_at 1439774_at 1458004_at | 8921 | 0.094 | -0.1461 | No | ||
| 58 | SMARCD2 | 1426192_at 1448400_a_at 1448401_at | 8952 | 0.092 | -0.1466 | No | ||
| 59 | EFNA3 | 1431170_at 1455344_at | 9160 | 0.080 | -0.1553 | No | ||
| 60 | PAX7 | 1444596_at 1452510_at | 9275 | 0.072 | -0.1599 | No | ||
| 61 | KCNAB1 | 1448468_a_at 1454043_a_at | 9305 | 0.070 | -0.1605 | No | ||
| 62 | CNKSR2 | 1444513_at 1445194_at | 9566 | 0.054 | -0.1720 | No | ||
| 63 | IL1F10 | 1451957_at | 9635 | 0.049 | -0.1746 | No | ||
| 64 | MOV10 | 1416380_at 1447376_at | 9762 | 0.042 | -0.1800 | No | ||
| 65 | PAK1IP1 | 1423766_at 1430875_a_at | 9886 | 0.033 | -0.1853 | No | ||
| 66 | RASL12 | 1428067_at | 10181 | 0.014 | -0.1987 | No | ||
| 67 | FXYD3 | 1418374_at | 10293 | 0.007 | -0.2038 | No | ||
| 68 | GSK3B | 1434439_at 1437001_at 1439931_at 1439949_at 1451020_at 1454958_at | 10461 | -0.003 | -0.2114 | No | ||
| 69 | CLIC5 | 1431261_at 1439505_at 1456873_at 1458381_at | 10487 | -0.005 | -0.2125 | No | ||
| 70 | NACA | 1416669_s_at | 10555 | -0.009 | -0.2155 | No | ||
| 71 | ARID1A | 1447104_at | 10559 | -0.009 | -0.2156 | No | ||
| 72 | ELF4 | 1421337_at 1445778_at | 10624 | -0.013 | -0.2184 | No | ||
| 73 | PTRF | 1421431_at 1424130_a_at 1453891_at | 10871 | -0.030 | -0.2294 | No | ||
| 74 | ROM1 | 1448996_at | 11050 | -0.042 | -0.2372 | No | ||
| 75 | BNC2 | 1438861_at 1440593_at 1459132_at | 11271 | -0.055 | -0.2468 | No | ||
| 76 | STOML2 | 1448774_at | 11585 | -0.073 | -0.2605 | No | ||
| 77 | MAZ | 1427099_at | 11658 | -0.078 | -0.2630 | No | ||
| 78 | BACE1 | 1421824_at 1421825_at 1436956_at 1450384_at 1455826_a_at | 11692 | -0.080 | -0.2638 | No | ||
| 79 | IFRD2 | 1451016_at | 11729 | -0.083 | -0.2646 | No | ||
| 80 | LIN28 | 1437752_at | 11747 | -0.084 | -0.2646 | No | ||
| 81 | TUBA3 | 1438611_at | 11794 | -0.086 | -0.2659 | No | ||
| 82 | BCL6 | 1421818_at 1450381_a_at | 11919 | -0.094 | -0.2707 | No | ||
| 83 | SALL1 | 1450489_at | 12122 | -0.105 | -0.2789 | No | ||
| 84 | CASQ2 | 1422529_s_at | 12177 | -0.109 | -0.2804 | No | ||
| 85 | CACNA1G | 1423365_at | 12295 | -0.116 | -0.2846 | No | ||
| 86 | TAL1 | 1449389_at | 12479 | -0.129 | -0.2918 | No | ||
| 87 | TRIP10 | 1418092_s_at | 12569 | -0.134 | -0.2946 | No | ||
| 88 | GABRG2 | 1418177_at 1437147_at | 12593 | -0.135 | -0.2944 | No | ||
| 89 | MYBPH | 1419487_at | 12623 | -0.137 | -0.2944 | No | ||
| 90 | WNT4 | 1450782_at | 12722 | -0.144 | -0.2975 | No | ||
| 91 | GABARAP | 1416937_at | 12739 | -0.145 | -0.2968 | No | ||
| 92 | RAB4B | 1451643_a_at | 13024 | -0.162 | -0.3083 | No | ||
| 93 | ARHGEF15 | 1440358_at 1455522_at 1457172_at | 13060 | -0.165 | -0.3084 | No | ||
| 94 | SEMA6A | 1421414_a_at 1425903_at 1436458_at | 13154 | -0.171 | -0.3110 | No | ||
| 95 | MRPL40 | 1448849_at | 13160 | -0.172 | -0.3096 | No | ||
| 96 | RHO | 1425171_at 1425172_at 1451617_at 1451618_at | 13263 | -0.178 | -0.3125 | No | ||
| 97 | SLC25A29 | 1423979_a_at 1423980_at 1423981_x_at 1438187_at 1438188_x_at | 13313 | -0.181 | -0.3130 | No | ||
| 98 | HARS2 | 1446462_at 1447677_x_at 1451040_at | 13417 | -0.189 | -0.3160 | No | ||
| 99 | OBSCN | 1443632_at | 13479 | -0.193 | -0.3169 | No | ||
| 100 | MYOG | 1419391_at | 13558 | -0.197 | -0.3186 | No | ||
| 101 | PCBP2 | 1421743_a_at 1435881_at | 13618 | -0.202 | -0.3194 | No | ||
| 102 | RFX4 | 1432053_at 1436931_at 1443312_at | 13660 | -0.205 | -0.3193 | No | ||
| 103 | MSX1 | 1417127_at 1448601_s_at | 13678 | -0.207 | -0.3181 | No | ||
| 104 | NDUFS2 | 1451096_at | 13769 | -0.212 | -0.3202 | No | ||
| 105 | PLA2G4D | 1453405_at | 13924 | -0.223 | -0.3251 | No | ||
| 106 | CRMP1 | 1448289_at | 13931 | -0.224 | -0.3232 | No | ||
| 107 | SLC7A10 | 1421093_at | 13969 | -0.226 | -0.3227 | No | ||
| 108 | AKT3 | 1422078_at 1435260_at 1435879_at 1443222_at 1460307_at | 14140 | -0.237 | -0.3283 | No | ||
| 109 | COX7A2L | 1421772_a_at 1432263_a_at 1432264_x_at | 14141 | -0.237 | -0.3260 | No | ||
| 110 | NKX2-3 | 1425827_at 1427741_x_at 1452528_a_at | 14259 | -0.246 | -0.3290 | No | ||
| 111 | EMILIN1 | 1416414_at 1431834_a_at | 14428 | -0.257 | -0.3342 | No | ||
| 112 | LRP8 | 1421459_a_at 1440882_at 1442347_at | 14618 | -0.273 | -0.3403 | No | ||
| 113 | OTOP2 | 1443043_at | 14649 | -0.275 | -0.3390 | No | ||
| 114 | MGEA5 | 1422900_at 1422901_at 1422902_s_at 1433654_at 1439081_at | 14763 | -0.283 | -0.3415 | No | ||
| 115 | LAG3 | 1449911_at | 14857 | -0.289 | -0.3430 | No | ||
| 116 | CIC | 1415746_at | 15350 | -0.325 | -0.3625 | No | ||
| 117 | ARID4A | 1436191_at 1438464_at 1456420_at | 15431 | -0.331 | -0.3630 | No | ||
| 118 | IMMT | 1429533_at 1429534_a_at 1433470_a_at 1437126_at | 15434 | -0.332 | -0.3599 | No | ||
| 119 | FKBP2 | 1450694_at | 15534 | -0.339 | -0.3612 | No | ||
| 120 | HOXC10 | 1439798_at | 15542 | -0.340 | -0.3582 | No | ||
| 121 | PRDM10 | 1441764_at 1444812_at | 15661 | -0.349 | -0.3603 | No | ||
| 122 | HEBP1 | 1418172_at 1457903_at | 15901 | -0.369 | -0.3677 | No | ||
| 123 | JPH4 | 1439031_at | 16204 | -0.394 | -0.3778 | No | ||
| 124 | TCF7 | 1433471_at 1450461_at | 16395 | -0.411 | -0.3826 | No | ||
| 125 | NFKBIA | 1420088_at 1420089_at 1438157_s_at 1448306_at 1449731_s_at | 16399 | -0.411 | -0.3788 | No | ||
| 126 | GPR4 | 1457745_at | 16443 | -0.416 | -0.3768 | No | ||
| 127 | PHF12 | 1434922_at 1453579_at | 16518 | -0.422 | -0.3761 | No | ||
| 128 | PPY | 1420440_at | 16545 | -0.425 | -0.3732 | No | ||
| 129 | HIRA | 1436241_s_at 1450049_a_at 1450050_at 1459103_at | 16624 | -0.431 | -0.3726 | No | ||
| 130 | CACNB1 | 1425777_at 1426108_s_at 1440940_at 1451834_at | 16725 | -0.439 | -0.3730 | No | ||
| 131 | ADAMTS4 | 1452595_at | 16792 | -0.447 | -0.3717 | No | ||
| 132 | LCAT | 1417043_at | 16960 | -0.464 | -0.3749 | No | ||
| 133 | DHH | 1422127_at 1434959_at | 17062 | -0.475 | -0.3750 | No | ||
| 134 | NPC2 | 1416901_at 1443266_at 1448513_a_at 1455768_at | 17101 | -0.479 | -0.3721 | No | ||
| 135 | PSD | 1435780_at | 17312 | -0.500 | -0.3770 | No | ||
| 136 | DLX1 | 1449470_at | 17325 | -0.502 | -0.3727 | No | ||
| 137 | RABGGTA | 1426046_a_at | 17462 | -0.518 | -0.3740 | No | ||
| 138 | CXCR4 | 1448710_at | 17598 | -0.531 | -0.3751 | No | ||
| 139 | RAB1B | 1424064_at | 17953 | -0.577 | -0.3858 | No | ||
| 140 | SOX5 | 1423500_a_at 1432189_a_at 1432190_at 1440827_x_at 1446461_at 1452511_at 1459261_at | 18107 | -0.598 | -0.3871 | No | ||
| 141 | PRKCBP1 | 1426614_at 1429415_at 1445827_at 1456143_at 1458617_at | 18328 | -0.627 | -0.3911 | Yes | ||
| 142 | LMOD3 | 1439658_at | 18434 | -0.642 | -0.3898 | Yes | ||
| 143 | SLC2A4 | 1415958_at 1415959_at | 18487 | -0.650 | -0.3859 | Yes | ||
| 144 | KLK9 | 1431681_at | 18517 | -0.656 | -0.3810 | Yes | ||
| 145 | CPNE1 | 1433439_at | 18590 | -0.666 | -0.3779 | Yes | ||
| 146 | NGFB | 1419675_at | 18653 | -0.675 | -0.3742 | Yes | ||
| 147 | THRAP1 | 1436904_at 1438725_at 1444068_at 1446840_at 1453160_at 1453161_at | 18741 | -0.691 | -0.3716 | Yes | ||
| 148 | ZFPM1 | 1423603_at 1451046_at | 18975 | -0.726 | -0.3753 | Yes | ||
| 149 | UPF2 | 1435030_at 1445777_at | 19004 | -0.730 | -0.3695 | Yes | ||
| 150 | NXF1 | 1416791_a_at | 19065 | -0.741 | -0.3652 | Yes | ||
| 151 | RHOG | 1422572_at | 19100 | -0.748 | -0.3595 | Yes | ||
| 152 | DCTN1 | 1422521_at 1435414_s_at | 19269 | -0.779 | -0.3598 | Yes | ||
| 153 | NCDN | 1416852_a_at 1416853_at | 19309 | -0.786 | -0.3540 | Yes | ||
| 154 | DDX17 | 1437773_x_at 1439037_at 1452155_a_at | 19502 | -0.822 | -0.3549 | Yes | ||
| 155 | ANXA4 | 1421223_a_at 1424176_a_at 1443341_at 1457658_x_at | 19503 | -0.822 | -0.3470 | Yes | ||
| 156 | GBA2 | 1419980_at 1434271_at | 19505 | -0.822 | -0.3391 | Yes | ||
| 157 | PYGO2 | 1428249_at | 19529 | -0.827 | -0.3322 | Yes | ||
| 158 | CREB3L2 | 1437396_at 1442080_at 1452381_at | 19842 | -0.894 | -0.3380 | Yes | ||
| 159 | MNT | 1418192_at | 19924 | -0.914 | -0.3329 | Yes | ||
| 160 | BRPF1 | 1460402_at | 20016 | -0.937 | -0.3280 | Yes | ||
| 161 | BOC | 1426869_at 1427516_a_at 1427517_at | 20210 | -0.984 | -0.3274 | Yes | ||
| 162 | DLGAP4 | 1426465_at 1455548_at | 20260 | -0.998 | -0.3201 | Yes | ||
| 163 | RASGRP2 | 1417804_at 1438932_at 1438933_x_at 1442264_at 1449613_at | 20346 | -1.013 | -0.3143 | Yes | ||
| 164 | TITF1 | 1422346_at | 20446 | -1.035 | -0.3088 | Yes | ||
| 165 | RBBP6 | 1425114_at 1425115_at 1425421_at 1426487_a_at 1453611_at | 20531 | -1.057 | -0.3025 | Yes | ||
| 166 | RBM12 | 1416928_at 1416929_at 1456964_at 1457934_at | 20550 | -1.064 | -0.2931 | Yes | ||
| 167 | POLD4 | 1427885_at | 20622 | -1.085 | -0.2859 | Yes | ||
| 168 | TUBA1 | 1418884_x_at | 20768 | -1.142 | -0.2816 | Yes | ||
| 169 | BAHD1 | 1433703_s_at | 20864 | -1.185 | -0.2746 | Yes | ||
| 170 | FLI1 | 1422024_at 1433512_at 1441584_at | 21162 | -1.346 | -0.2753 | Yes | ||
| 171 | ZFP36 | 1452519_a_at | 21206 | -1.378 | -0.2640 | Yes | ||
| 172 | SH3BP1 | 1460222_at | 21248 | -1.417 | -0.2522 | Yes | ||
| 173 | TUBA4 | 1417373_a_at 1417374_at 1417375_at | 21293 | -1.454 | -0.2402 | Yes | ||
| 174 | PLEC1 | 1419835_s_at 1434610_at 1437554_at | 21346 | -1.505 | -0.2282 | Yes | ||
| 175 | NRGN | 1423231_at | 21361 | -1.514 | -0.2142 | Yes | ||
| 176 | GNAO1 | 1421152_a_at 1448031_at 1460383_at | 21374 | -1.524 | -0.2001 | Yes | ||
| 177 | DMPK | 1422989_a_at 1434944_at | 21442 | -1.597 | -0.1878 | Yes | ||
| 178 | ETV1 | 1422607_at 1445314_at 1450684_at | 21566 | -1.747 | -0.1766 | Yes | ||
| 179 | DPF2 | 1416534_at 1429903_at | 21596 | -1.808 | -0.1606 | Yes | ||
| 180 | VASP | 1451097_at | 21643 | -1.914 | -0.1443 | Yes | ||
| 181 | EDG1 | 1423571_at | 21820 | -2.744 | -0.1259 | Yes | ||
| 182 | SESN3 | 1449303_at 1453313_at | 21827 | -2.823 | -0.0990 | Yes | ||
| 183 | CNN2 | 1450981_at | 21853 | -3.054 | -0.0708 | Yes | ||
| 184 | CKB | 1455106_a_at | 21906 | -3.736 | -0.0372 | Yes | ||
| 185 | NR4A1 | 1416505_at | 21917 | -3.996 | 0.0008 | Yes |