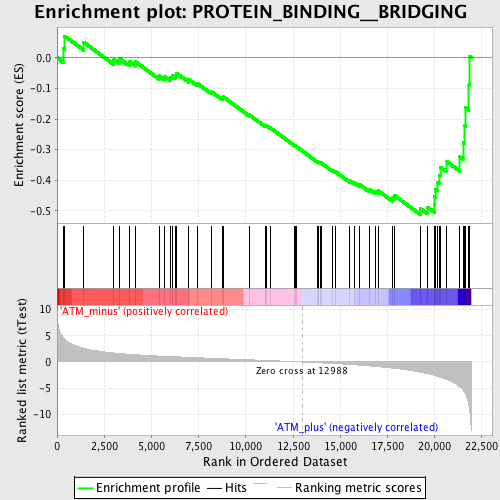
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | PROTEIN_BINDING__BRIDGING |
| Enrichment Score (ES) | -0.51143944 |
| Normalized Enrichment Score (NES) | -1.6929055 |
| Nominal p-value | 0.0016207455 |
| FDR q-value | 0.2429815 |
| FWER p-Value | 0.993 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CRKL | 1421953_at 1421954_at 1425604_at 1436950_at | 324 | 4.648 | 0.0314 | No | ||
| 2 | SHB | 1434153_at | 406 | 4.319 | 0.0706 | No | ||
| 3 | LOR | 1420183_at 1448745_s_at | 1415 | 2.604 | 0.0505 | No | ||
| 4 | ANXA1 | 1448213_at | 2994 | 1.721 | -0.0046 | No | ||
| 5 | GRB14 | 1417673_at 1457526_at | 3285 | 1.626 | -0.0016 | No | ||
| 6 | ABI2 | 1433985_at 1436984_at 1444783_at 1458053_at 1460153_at | 3825 | 1.468 | -0.0117 | No | ||
| 7 | STUB1 | 1416580_a_at 1429069_at 1434412_x_at | 4160 | 1.380 | -0.0132 | No | ||
| 8 | BLNK | 1451780_at | 5398 | 1.127 | -0.0586 | No | ||
| 9 | ARHGAP6 | 1417704_a_at 1451867_x_at 1456333_a_at | 5706 | 1.071 | -0.0619 | No | ||
| 10 | CNTNAP1 | 1421580_at | 5982 | 1.029 | -0.0643 | No | ||
| 11 | SKAP1 | 1437249_at 1440326_at 1440782_at 1441322_at 1441418_at 1441589_at 1456678_at | 6086 | 1.012 | -0.0589 | No | ||
| 12 | IRS4 | 1441429_at | 6293 | 0.983 | -0.0586 | No | ||
| 13 | ARHGAP4 | 1419296_at 1443360_x_at 1458190_at | 6334 | 0.976 | -0.0507 | No | ||
| 14 | LASP1 | 1438633_x_at 1438634_x_at 1439264_x_at 1444947_at 1448207_at 1455470_x_at 1456309_x_at 1456469_x_at 1456578_x_at 1460173_at | 6972 | 0.881 | -0.0710 | No | ||
| 15 | GRB2 | 1418508_a_at 1449111_a_at | 7424 | 0.806 | -0.0836 | No | ||
| 16 | SPRR1B | 1422672_at | 8182 | 0.696 | -0.1113 | No | ||
| 17 | EPS8 | 1422823_at 1422824_s_at 1425733_a_at 1436977_at | 8742 | 0.613 | -0.1308 | No | ||
| 18 | DSP | 1427610_at 1435493_at 1435494_s_at | 8801 | 0.604 | -0.1274 | No | ||
| 19 | ITSN2 | 1423184_at 1431772_a_at 1435023_at 1445375_at 1446595_at 1446735_at 1446911_at | 10187 | 0.413 | -0.1866 | No | ||
| 20 | SNX9 | 1423076_at 1423077_at | 11023 | 0.298 | -0.2218 | No | ||
| 21 | SH3BGR | 1422644_at | 11090 | 0.288 | -0.2220 | No | ||
| 22 | SH3BGRL | 1421871_at 1428107_at 1436997_x_at 1459571_at | 11281 | 0.258 | -0.2281 | No | ||
| 23 | RAD50 | 1422630_at 1425939_at 1458970_at | 12597 | 0.064 | -0.2875 | No | ||
| 24 | EVPL | 1419133_at | 12602 | 0.063 | -0.2871 | No | ||
| 25 | GRB7 | 1448227_at | 12679 | 0.051 | -0.2901 | No | ||
| 26 | LAT | 1460651_at | 13781 | -0.141 | -0.3390 | No | ||
| 27 | GRB10 | 1425457_a_at 1425458_a_at 1430164_a_at 1440935_at 1441620_at 1446921_at | 13858 | -0.153 | -0.3409 | No | ||
| 28 | CHN1 | 1420545_a_at 1445691_at | 13946 | -0.168 | -0.3433 | No | ||
| 29 | FKBP4 | 1416362_a_at 1416363_at 1458729_at 1459808_at | 13985 | -0.176 | -0.3432 | No | ||
| 30 | CSTA | 1435760_at | 14569 | -0.281 | -0.3671 | No | ||
| 31 | SLA2 | 1437504_at 1451492_at | 14749 | -0.318 | -0.3721 | No | ||
| 32 | SKAP2 | 1418895_at 1457097_at 1460623_at | 15490 | -0.478 | -0.4012 | No | ||
| 33 | KHDRBS1 | 1418628_at 1418630_at | 15742 | -0.545 | -0.4072 | No | ||
| 34 | SPRR1A | 1449133_at | 16025 | -0.607 | -0.4141 | No | ||
| 35 | VAV3 | 1417122_at 1417123_at 1446795_at 1448600_s_at 1458630_at | 16529 | -0.754 | -0.4296 | No | ||
| 36 | ARPC4 | 1423588_at 1423589_at | 16835 | -0.852 | -0.4351 | No | ||
| 37 | SRC | 1423240_at 1450918_s_at | 16997 | -0.899 | -0.4335 | No | ||
| 38 | IVL | 1422222_at 1439878_at | 17759 | -1.178 | -0.4566 | No | ||
| 39 | RUSC1 | 1431137_at 1434743_x_at 1436014_a_at 1437991_x_at 1438017_at | 17870 | -1.223 | -0.4494 | No | ||
| 40 | CHN2 | 1428573_at 1428574_a_at 1441158_at 1443626_at | 19228 | -1.965 | -0.4919 | Yes | ||
| 41 | GRAP2 | 1456432_at | 19612 | -2.207 | -0.4875 | Yes | ||
| 42 | GAB1 | 1417693_a_at 1417694_at 1448814_at | 19983 | -2.564 | -0.4789 | Yes | ||
| 43 | ARHGAP1 | 1424307_at 1451309_at | 19996 | -2.579 | -0.4538 | Yes | ||
| 44 | GATAD2A | 1423992_at 1445239_at 1451197_s_at 1451198_at 1455505_at | 20042 | -2.617 | -0.4298 | Yes | ||
| 45 | SLA | 1420818_at 1420819_at 1441761_at 1447813_x_at | 20152 | -2.752 | -0.4075 | Yes | ||
| 46 | SH2D2A | 1449105_at | 20248 | -2.853 | -0.3834 | Yes | ||
| 47 | SOCS2 | 1418507_s_at 1438470_at 1441476_at 1442586_at 1446085_at 1449109_at | 20296 | -2.903 | -0.3567 | Yes | ||
| 48 | BCL3 | 1418133_at | 20646 | -3.380 | -0.3391 | Yes | ||
| 49 | STAM | 1416861_at 1416862_at 1457828_at 1459427_at | 21320 | -4.700 | -0.3231 | Yes | ||
| 50 | SH2D3C | 1415886_at | 21530 | -5.528 | -0.2777 | Yes | ||
| 51 | TOB1 | 1423176_at 1440844_at | 21595 | -5.891 | -0.2221 | Yes | ||
| 52 | SH3BP2 | 1448328_at | 21639 | -6.137 | -0.1630 | Yes | ||
| 53 | GRAP | 1429387_at | 21812 | -8.416 | -0.0872 | Yes | ||
| 54 | ST13 | 1416851_at 1442775_at 1460193_at | 21858 | -9.331 | 0.0035 | Yes |

