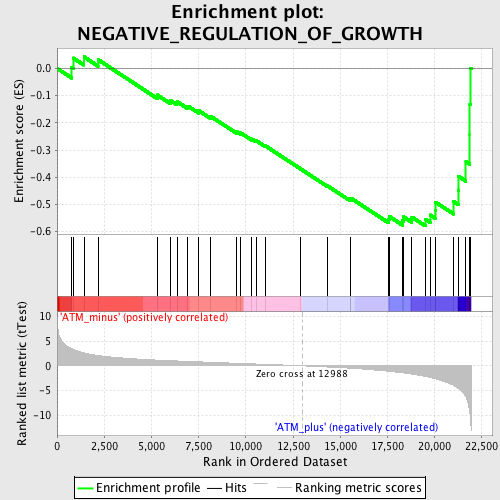
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | NEGATIVE_REGULATION_OF_GROWTH |
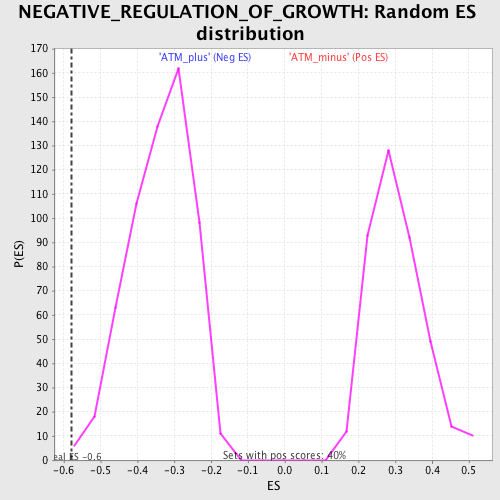
| Enrichment Score (ES) | -0.57887805 |
| Normalized Enrichment Score (NES) | -1.7201604 |
| Nominal p-value | 0.003322259 |
| FDR q-value | 0.33075008 |
| FWER p-Value | 0.968 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDKN1A | 1421679_a_at 1424638_at | 774 | 3.430 | 0.0043 | No | ||
| 2 | TGFB1 | 1420653_at 1445360_at | 856 | 3.267 | 0.0384 | No | ||
| 3 | CDKN2A | 1450140_a_at | 1423 | 2.597 | 0.0426 | No | ||
| 4 | NDUFS3 | 1423737_at | 2167 | 2.064 | 0.0325 | No | ||
| 5 | ENO1 | 1419023_x_at | 5302 | 1.142 | -0.0974 | No | ||
| 6 | TSPYL2 | 1434849_at | 6007 | 1.025 | -0.1177 | No | ||
| 7 | APBB1 | 1423892_at 1423893_x_at | 6350 | 0.974 | -0.1220 | No | ||
| 8 | APBB2 | 1426719_at 1426720_at 1440135_at 1446481_at 1452342_at | 6929 | 0.888 | -0.1382 | No | ||
| 9 | TP53 | 1426538_a_at 1427739_a_at 1438808_at 1457623_x_at 1459780_at 1459781_x_at | 7499 | 0.796 | -0.1549 | No | ||
| 10 | BMP10 | 1421763_at | 8132 | 0.704 | -0.1757 | No | ||
| 11 | CDKN2C | 1416868_at 1439164_at | 9503 | 0.503 | -0.2324 | No | ||
| 12 | CDA | 1427357_at | 9701 | 0.477 | -0.2359 | No | ||
| 13 | PPP1R9B | 1447247_at 1447486_at | 10311 | 0.396 | -0.2591 | No | ||
| 14 | SMAD3 | 1450471_at 1450472_s_at 1454960_at | 10548 | 0.360 | -0.2657 | No | ||
| 15 | PPT1 | 1420015_s_at 1420016_at 1422467_at 1422468_at 1444884_at | 11039 | 0.295 | -0.2847 | No | ||
| 16 | RERG | 1447446_at 1451236_at | 12879 | 0.021 | -0.3684 | No | ||
| 17 | DLC1 | 1436173_at 1450206_at 1458220_at 1460602_at | 14319 | -0.238 | -0.4314 | No | ||
| 18 | RB1 | 1417850_at 1441494_at 1444400_at 1444454_at 1457447_at | 15532 | -0.490 | -0.4810 | No | ||
| 19 | ING4 | 1448054_at 1460728_s_at | 15551 | -0.496 | -0.4761 | No | ||
| 20 | PTCH1 | 1428853_at 1439663_at 1447039_at 1450824_at | 17546 | -1.084 | -0.5547 | No | ||
| 21 | DCBLD2 | 1420526_at 1437635_at 1449891_a_at | 17589 | -1.100 | -0.5438 | No | ||
| 22 | INHBA | 1422053_at 1458291_at | 18304 | -1.420 | -0.5600 | Yes | ||
| 23 | PML | 1448757_at 1456103_at 1459137_at | 18354 | -1.441 | -0.5456 | Yes | ||
| 24 | ING5 | 1431134_at 1436069_at 1442094_at 1454235_a_at | 18788 | -1.664 | -0.5461 | Yes | ||
| 25 | IHPK2 | 1428373_at 1435319_at | 19506 | -2.133 | -0.5542 | Yes | ||
| 26 | SERTAD2 | 1417209_at 1444645_at 1454815_at | 19753 | -2.334 | -0.5385 | Yes | ||
| 27 | SMAD4 | 1422485_at 1422486_a_at 1422487_at 1444205_at | 20022 | -2.598 | -0.5207 | Yes | ||
| 28 | NDUFA13 | 1430713_s_at | 20037 | -2.613 | -0.4911 | Yes | ||
| 29 | CDKN1B | 1419497_at 1434045_at | 21001 | -3.924 | -0.4897 | Yes | ||
| 30 | SERTAD3 | 1421076_at 1421077_at | 21241 | -4.504 | -0.4485 | Yes | ||
| 31 | BBC3 | 1423315_at | 21263 | -4.564 | -0.3967 | Yes | ||
| 32 | ACVR1B | 1422098_at 1433725_at | 21652 | -6.207 | -0.3426 | Yes | ||
| 33 | KLK8 | 1419722_at | 21860 | -9.470 | -0.2426 | Yes | ||
| 34 | CDKN2D | 1416253_at | 21862 | -9.545 | -0.1323 | Yes | ||
| 35 | BCL6 | 1421818_at 1450381_a_at | 21907 | -11.718 | 0.0012 | Yes |