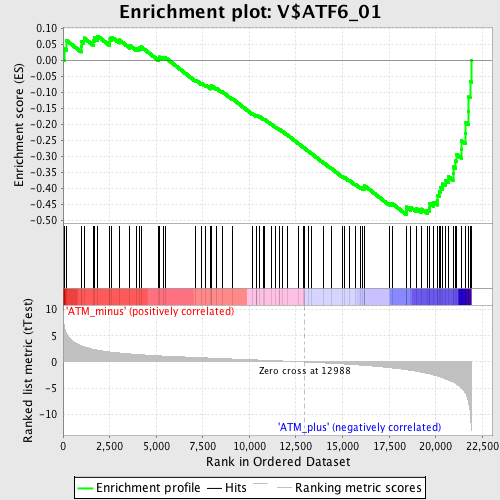
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | V$ATF6_01 |
| Enrichment Score (ES) | -0.48097405 |
| Normalized Enrichment Score (NES) | -1.7247043 |
| Nominal p-value | 0.0014947683 |
| FDR q-value | 0.022195024 |
| FWER p-Value | 0.218 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDELR2 | 1417204_at 1417205_at | 51 | 7.145 | 0.0376 | No | ||
| 2 | MRPS18B | 1451164_a_at | 188 | 5.327 | 0.0611 | No | ||
| 3 | GFRA1 | 1421973_at 1439015_at 1450440_at | 968 | 3.098 | 0.0427 | No | ||
| 4 | KDELR3 | 1418538_at | 997 | 3.060 | 0.0585 | No | ||
| 5 | TPM4 | 1433883_at | 1121 | 2.913 | 0.0691 | No | ||
| 6 | ERF | 1422114_at 1435561_at | 1612 | 2.417 | 0.0602 | No | ||
| 7 | SRP19 | 1450891_at | 1677 | 2.374 | 0.0705 | No | ||
| 8 | EIF4A1 | 1427058_at 1430980_a_at 1434985_a_at | 1856 | 2.244 | 0.0749 | No | ||
| 9 | SYNCRIP | 1422768_at 1422769_at 1426402_at 1428546_at 1450743_s_at | 2498 | 1.902 | 0.0562 | No | ||
| 10 | TWIST1 | 1418733_at | 2515 | 1.898 | 0.0660 | No | ||
| 11 | ADAMTS3 | 1441693_at 1458143_at | 2615 | 1.856 | 0.0719 | No | ||
| 12 | PTOV1 | 1416945_at 1438058_s_at 1458357_x_at | 3019 | 1.710 | 0.0630 | No | ||
| 13 | SEMA3B | 1416644_a_at 1431795_a_at 1448415_a_at | 3581 | 1.533 | 0.0459 | No | ||
| 14 | FLNC | 1424063_at 1447812_x_at 1449073_at | 3937 | 1.437 | 0.0376 | No | ||
| 15 | SEC24D | 1426972_at 1458821_at | 4078 | 1.399 | 0.0390 | No | ||
| 16 | USP2 | 1417168_a_at 1417169_at | 4185 | 1.375 | 0.0418 | No | ||
| 17 | NANS | 1417773_at 1417774_at | 5111 | 1.180 | 0.0061 | No | ||
| 18 | TRPC1 | 1421095_a_at 1421096_at | 5170 | 1.170 | 0.0100 | No | ||
| 19 | LHX5 | 1420348_at | 5366 | 1.132 | 0.0074 | No | ||
| 20 | CREM | 1418322_at 1430598_at 1430847_a_at 1449037_at | 5491 | 1.109 | 0.0079 | No | ||
| 21 | LENG9 | 1440190_at 1440191_s_at 1455497_at | 7134 | 0.852 | -0.0625 | No | ||
| 22 | GRB2 | 1418508_a_at 1449111_a_at | 7424 | 0.806 | -0.0713 | No | ||
| 23 | SLC39A5 | 1429523_a_at | 7659 | 0.772 | -0.0777 | No | ||
| 24 | NOL4 | 1430189_at 1441447_at 1456387_at | 7928 | 0.736 | -0.0858 | No | ||
| 25 | HOXB9 | 1452317_at | 7957 | 0.732 | -0.0830 | No | ||
| 26 | ARF4 | 1423052_at 1423053_at 1434074_x_at 1439367_x_at 1442905_at 1457348_at | 7971 | 0.731 | -0.0795 | No | ||
| 27 | LMX1B | 1421769_at | 8218 | 0.690 | -0.0869 | No | ||
| 28 | NKX2-2 | 1421112_at | 8544 | 0.641 | -0.0982 | No | ||
| 29 | GRIA3 | 1420563_at 1434728_at | 9071 | 0.566 | -0.1192 | No | ||
| 30 | XPNPEP1 | 1422443_at | 10150 | 0.417 | -0.1662 | No | ||
| 31 | SLC22A17 | 1448209_a_at | 10370 | 0.387 | -0.1740 | No | ||
| 32 | HCRTR2 | 1444223_at | 10379 | 0.386 | -0.1722 | No | ||
| 33 | GSC | 1421412_at | 10405 | 0.381 | -0.1713 | No | ||
| 34 | DHX40 | 1428267_at 1457839_at | 10551 | 0.360 | -0.1759 | No | ||
| 35 | ETV6 | 1423401_at 1434880_at 1444598_at 1457944_at | 10774 | 0.329 | -0.1842 | No | ||
| 36 | GDNF | 1419080_at | 10801 | 0.326 | -0.1836 | No | ||
| 37 | SLC30A5 | 1422497_at | 11179 | 0.273 | -0.1993 | No | ||
| 38 | SYT11 | 1429314_at 1429315_at 1429729_at 1436176_at 1449264_at 1455176_a_at | 11410 | 0.239 | -0.2085 | No | ||
| 39 | NTRK2 | 1420837_at 1420838_at 1435196_at 1435305_at 1437560_at 1446712_at | 11623 | 0.209 | -0.2171 | No | ||
| 40 | STAG1 | 1421939_a_at 1421940_at 1431921_a_at 1434189_at 1435731_x_at 1445145_at 1445652_at 1450420_at | 11636 | 0.207 | -0.2164 | No | ||
| 41 | GOLGB1 | 1433135_at 1435032_at 1460535_at | 11755 | 0.189 | -0.2208 | No | ||
| 42 | PITX2 | 1424797_a_at 1450482_a_at | 12026 | 0.147 | -0.2323 | No | ||
| 43 | TBX20 | 1422219_a_at 1425158_at 1453351_at | 12651 | 0.056 | -0.2606 | No | ||
| 44 | NPTX1 | 1422130_at 1434877_at | 12911 | 0.014 | -0.2724 | No | ||
| 45 | IL1RAPL1 | 1445821_at 1458964_at | 12936 | 0.010 | -0.2734 | No | ||
| 46 | DLST | 1423710_at 1430161_at 1431011_at 1437775_at | 12974 | 0.002 | -0.2751 | No | ||
| 47 | DUSP4 | 1428834_at | 13169 | -0.032 | -0.2838 | No | ||
| 48 | TPT1 | 1416642_a_at 1416643_at | 13338 | -0.060 | -0.2911 | No | ||
| 49 | PAX1 | 1449359_at | 13997 | -0.178 | -0.3203 | No | ||
| 50 | IRX3 | 1418517_at | 14385 | -0.248 | -0.3366 | No | ||
| 51 | GRIN2B | 1422223_at 1431700_at | 15000 | -0.370 | -0.3626 | No | ||
| 52 | KCNA6 | 1421568_at 1441049_at 1456954_at | 15118 | -0.390 | -0.3658 | No | ||
| 53 | TYRO3 | 1425248_a_at 1425249_a_at | 15357 | -0.445 | -0.3742 | No | ||
| 54 | ICA1 | 1417901_a_at 1431644_a_at | 15685 | -0.532 | -0.3862 | No | ||
| 55 | EGR3 | 1421486_at 1436329_at | 15964 | -0.593 | -0.3957 | No | ||
| 56 | TLOC1 | 1432635_a_at 1433704_s_at 1439233_at 1444811_at 1454697_at 1455698_at | 16103 | -0.631 | -0.3984 | No | ||
| 57 | TLE3 | 1419654_at 1419655_at 1449554_at 1458512_at | 16179 | -0.650 | -0.3983 | No | ||
| 58 | TOP1 | 1423474_at 1438495_at 1457573_at 1459672_at | 16190 | -0.655 | -0.3950 | No | ||
| 59 | LHX1 | 1421951_at 1450428_at | 16193 | -0.657 | -0.3915 | No | ||
| 60 | TNKS2 | 1428388_at 1437793_at 1447522_s_at 1452772_at 1457468_at | 17507 | -1.068 | -0.4456 | No | ||
| 61 | LMO4 | 1420981_a_at 1439651_at | 17681 | -1.145 | -0.4471 | No | ||
| 62 | C1QTNF7 | 1429030_at 1460163_at | 18421 | -1.474 | -0.4727 | Yes | ||
| 63 | RAB2 | 1418621_at 1418622_at 1418623_at 1419945_s_at 1419946_s_at 1436477_x_at 1449676_at | 18438 | -1.481 | -0.4652 | Yes | ||
| 64 | SEC23A | 1423347_at | 18452 | -1.487 | -0.4575 | Yes | ||
| 65 | SFRS1 | 1428099_a_at 1428100_at 1430982_at 1434972_x_at 1452430_s_at 1453722_s_at 1457136_at | 18680 | -1.587 | -0.4590 | Yes | ||
| 66 | KDELR1 | 1460211_a_at | 18981 | -1.783 | -0.4628 | Yes | ||
| 67 | GNG4 | 1417943_at 1417944_at 1447669_s_at | 19246 | -1.976 | -0.4639 | Yes | ||
| 68 | OSBP | 1460350_at | 19582 | -2.186 | -0.4670 | Yes | ||
| 69 | SLC39A13 | 1423926_at | 19657 | -2.247 | -0.4578 | Yes | ||
| 70 | KLF13 | 1432543_a_at 1456030_at | 19698 | -2.278 | -0.4470 | Yes | ||
| 71 | PPP1R10 | 1426726_at 1426727_s_at 1430560_at | 19898 | -2.477 | -0.4422 | Yes | ||
| 72 | SCYL1 | 1430354_x_at 1436804_s_at 1451051_a_at | 20102 | -2.698 | -0.4365 | Yes | ||
| 73 | BRD2 | 1423502_at 1437210_a_at | 20113 | -2.714 | -0.4218 | Yes | ||
| 74 | PER1 | 1449851_at | 20193 | -2.792 | -0.4098 | Yes | ||
| 75 | SH2D2A | 1449105_at | 20248 | -2.853 | -0.3964 | Yes | ||
| 76 | ATBF1 | 1420649_at 1420650_at 1429725_at 1445941_at 1446094_at 1449947_s_at 1453267_at 1460006_at | 20385 | -3.010 | -0.3858 | Yes | ||
| 77 | PHC1 | 1423212_at | 20522 | -3.189 | -0.3742 | Yes | ||
| 78 | STAT3 | 1424272_at 1426587_a_at 1459961_a_at 1460700_at | 20682 | -3.421 | -0.3624 | Yes | ||
| 79 | GCC1 | 1424799_a_at 1429033_at | 20942 | -3.816 | -0.3529 | Yes | ||
| 80 | SP1 | 1418180_at 1448994_at 1454852_at | 20975 | -3.869 | -0.3328 | Yes | ||
| 81 | ELF1 | 1417540_at 1439319_at | 21063 | -4.056 | -0.3141 | Yes | ||
| 82 | TRIM39 | 1418851_at | 21110 | -4.153 | -0.2931 | Yes | ||
| 83 | GBF1 | 1415709_s_at 1415757_at 1432677_at 1438207_at | 21378 | -4.867 | -0.2781 | Yes | ||
| 84 | ARMCX2 | 1415780_a_at 1437849_x_at 1456739_x_at | 21386 | -4.885 | -0.2512 | Yes | ||
| 85 | RORC | 1425792_a_at 1425793_a_at | 21592 | -5.874 | -0.2277 | Yes | ||
| 86 | MADD | 1425735_at 1455502_at | 21632 | -6.088 | -0.1955 | Yes | ||
| 87 | NR4A1 | 1416505_at | 21771 | -7.733 | -0.1587 | Yes | ||
| 88 | RAB6IP1 | 1424015_at | 21789 | -8.026 | -0.1147 | Yes | ||
| 89 | EGR1 | 1417065_at | 21855 | -9.280 | -0.0658 | Yes | ||
| 90 | EGR2 | 1427682_a_at 1427683_at | 21928 | -12.431 | 0.0003 | Yes |

