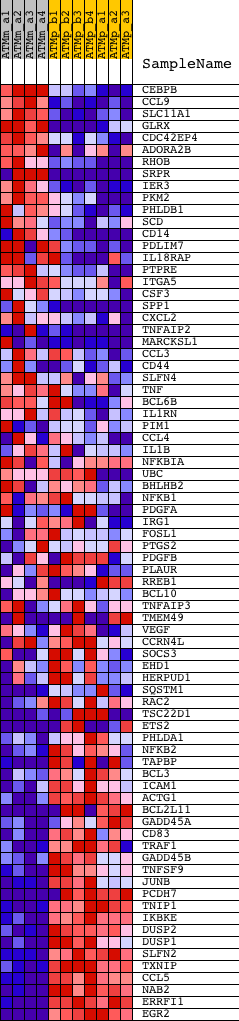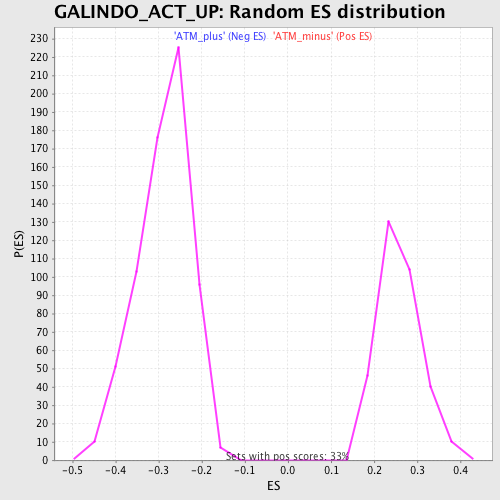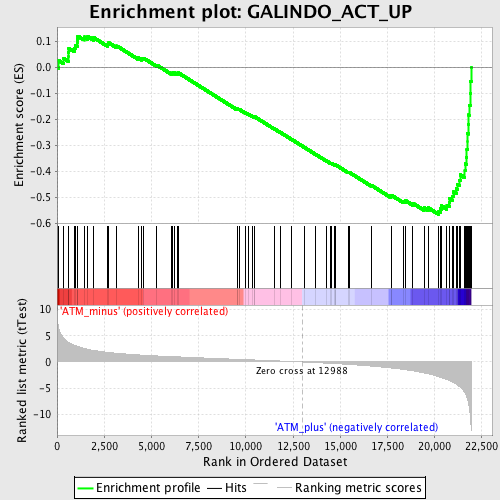
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | GALINDO_ACT_UP |
| Enrichment Score (ES) | -0.56654793 |
| Normalized Enrichment Score (NES) | -1.9770217 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.013512397 |
| FWER p-Value | 0.102 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CEBPB | 1418901_at 1427844_a_at | 86 | 6.374 | 0.0249 | No | ||
| 2 | CCL9 | 1417936_at 1448898_at | 354 | 4.559 | 0.0333 | No | ||
| 3 | SLC11A1 | 1420361_at | 588 | 3.777 | 0.0397 | No | ||
| 4 | GLRX | 1416592_at 1416593_at | 593 | 3.763 | 0.0566 | No | ||
| 5 | CDC42EP4 | 1416511_a_at 1436801_x_at | 621 | 3.701 | 0.0721 | No | ||
| 6 | ADORA2B | 1434430_s_at 1434431_x_at 1434772_at 1450214_at | 908 | 3.182 | 0.0734 | No | ||
| 7 | RHOB | 1449110_at | 974 | 3.091 | 0.0844 | No | ||
| 8 | SRPR | 1423670_a_at 1427570_at | 1067 | 2.979 | 0.0936 | No | ||
| 9 | IER3 | 1419647_a_at | 1077 | 2.972 | 0.1067 | No | ||
| 10 | PKM2 | 1417308_at | 1097 | 2.950 | 0.1191 | No | ||
| 11 | PHLDB1 | 1424467_at 1424468_s_at | 1425 | 2.593 | 0.1159 | No | ||
| 12 | SCD | 1415964_at 1415965_at | 1628 | 2.405 | 0.1175 | No | ||
| 13 | CD14 | 1417268_at | 1943 | 2.195 | 0.1131 | No | ||
| 14 | PDLIM7 | 1417959_at 1428319_at 1443785_x_at | 2659 | 1.838 | 0.0887 | No | ||
| 15 | IL18RAP | 1421291_at 1456545_at | 2741 | 1.810 | 0.0932 | No | ||
| 16 | PTPRE | 1418539_a_at 1418540_a_at 1459638_at | 3166 | 1.659 | 0.0813 | No | ||
| 17 | ITGA5 | 1423267_s_at 1423268_at 1457561_at 1458996_at | 4293 | 1.353 | 0.0359 | No | ||
| 18 | CSF3 | 1419427_at | 4491 | 1.302 | 0.0328 | No | ||
| 19 | SPP1 | 1449254_at | 4600 | 1.277 | 0.0336 | No | ||
| 20 | CXCL2 | 1449984_at | 5274 | 1.148 | 0.0080 | No | ||
| 21 | TNFAIP2 | 1416273_at 1438855_x_at 1457918_at | 6048 | 1.019 | -0.0228 | No | ||
| 22 | MARCKSL1 | 1415922_s_at 1435415_x_at 1437226_x_at | 6132 | 1.006 | -0.0220 | No | ||
| 23 | CCL3 | 1419561_at | 6237 | 0.991 | -0.0223 | No | ||
| 24 | CD44 | 1423760_at 1434376_at 1443265_at 1452483_a_at | 6384 | 0.968 | -0.0246 | No | ||
| 25 | SLFN4 | 1427102_at | 6423 | 0.963 | -0.0220 | No | ||
| 26 | TNF | 1419607_at | 9533 | 0.499 | -0.1620 | No | ||
| 27 | BCL6B | 1418421_at | 9540 | 0.498 | -0.1600 | No | ||
| 28 | IL1RN | 1423017_a_at 1425663_at 1451798_at | 9632 | 0.486 | -0.1619 | No | ||
| 29 | PIM1 | 1423006_at 1435458_at | 9985 | 0.437 | -0.1761 | No | ||
| 30 | CCL4 | 1421578_at | 10152 | 0.417 | -0.1818 | No | ||
| 31 | IL1B | 1449399_a_at | 10345 | 0.391 | -0.1888 | No | ||
| 32 | NFKBIA | 1420088_at 1420089_at 1438157_s_at 1448306_at 1449731_s_at | 10443 | 0.375 | -0.1915 | No | ||
| 33 | UBC | 1420494_x_at 1425965_at 1425966_x_at 1432827_x_at 1437666_x_at 1454373_x_at | 10479 | 0.370 | -0.1915 | No | ||
| 34 | BHLHB2 | 1418025_at | 11517 | 0.223 | -0.2379 | No | ||
| 35 | NFKB1 | 1427705_a_at 1442949_at | 11848 | 0.175 | -0.2522 | No | ||
| 36 | PDGFA | 1418711_at 1449187_at | 12405 | 0.091 | -0.2772 | No | ||
| 37 | IRG1 | 1427381_at | 13123 | -0.024 | -0.3099 | No | ||
| 38 | FOSL1 | 1417487_at 1417488_at | 13667 | -0.117 | -0.3342 | No | ||
| 39 | PTGS2 | 1417262_at 1417263_at | 14273 | -0.228 | -0.3609 | No | ||
| 40 | PDGFB | 1450413_at 1450414_at | 14499 | -0.271 | -0.3700 | No | ||
| 41 | PLAUR | 1452521_a_at | 14509 | -0.272 | -0.3691 | No | ||
| 42 | RREB1 | 1428657_at 1434741_at 1438216_at 1442047_at 1452862_at | 14699 | -0.308 | -0.3764 | No | ||
| 43 | BCL10 | 1418970_a_at 1418971_x_at 1418972_at 1443524_x_at | 14734 | -0.314 | -0.3765 | No | ||
| 44 | TNFAIP3 | 1433699_at 1450829_at | 14751 | -0.319 | -0.3758 | No | ||
| 45 | TMEM49 | 1421491_a_at 1423722_at 1444820_at | 15408 | -0.457 | -0.4038 | No | ||
| 46 | VEGF | 1420909_at 1451959_a_at | 15473 | -0.473 | -0.4046 | No | ||
| 47 | CCRN4L | 1425837_a_at 1458699_at | 16647 | -0.795 | -0.4546 | No | ||
| 48 | SOCS3 | 1416576_at 1455899_x_at 1456212_x_at | 17683 | -1.146 | -0.4968 | No | ||
| 49 | EHD1 | 1416010_a_at 1416011_x_at 1416012_at 1448175_at | 17710 | -1.159 | -0.4927 | No | ||
| 50 | HERPUD1 | 1435626_a_at 1448185_at | 18341 | -1.435 | -0.5151 | No | ||
| 51 | SQSTM1 | 1440076_at 1442237_at 1444021_at 1450957_a_at | 18467 | -1.501 | -0.5140 | No | ||
| 52 | RAC2 | 1417620_at 1440208_at | 18839 | -1.688 | -0.5233 | No | ||
| 53 | TSC22D1 | 1425742_a_at 1433899_x_at 1447360_at 1454758_a_at 1454971_x_at 1456132_x_at | 19434 | -2.093 | -0.5411 | No | ||
| 54 | ETS2 | 1416268_at 1437809_x_at 1447685_x_at | 19651 | -2.244 | -0.5408 | No | ||
| 55 | PHLDA1 | 1418835_at | 20215 | -2.819 | -0.5538 | Yes | ||
| 56 | NFKB2 | 1425902_a_at 1429128_x_at | 20307 | -2.917 | -0.5448 | Yes | ||
| 57 | TAPBP | 1421812_at 1450378_at | 20352 | -2.979 | -0.5333 | Yes | ||
| 58 | BCL3 | 1418133_at | 20646 | -3.380 | -0.5314 | Yes | ||
| 59 | ICAM1 | 1424067_at | 20776 | -3.566 | -0.5212 | Yes | ||
| 60 | ACTG1 | 1415779_s_at | 20795 | -3.592 | -0.5058 | Yes | ||
| 61 | BCL2L11 | 1426334_a_at 1435448_at 1435449_at 1456005_a_at 1456006_at 1459794_at | 20953 | -3.830 | -0.4956 | Yes | ||
| 62 | GADD45A | 1449519_at | 20994 | -3.913 | -0.4798 | Yes | ||
| 63 | CD83 | 1416111_at | 21143 | -4.248 | -0.4673 | Yes | ||
| 64 | TRAF1 | 1423602_at 1445452_at | 21192 | -4.381 | -0.4497 | Yes | ||
| 65 | GADD45B | 1420197_at 1449773_s_at 1450971_at | 21323 | -4.709 | -0.4344 | Yes | ||
| 66 | TNFSF9 | 1422924_at | 21349 | -4.785 | -0.4139 | Yes | ||
| 67 | JUNB | 1415899_at | 21600 | -5.913 | -0.3986 | Yes | ||
| 68 | PCDH7 | 1427592_at 1437442_at 1449249_at 1456214_at | 21609 | -5.972 | -0.3719 | Yes | ||
| 69 | TNIP1 | 1427689_a_at | 21690 | -6.558 | -0.3459 | Yes | ||
| 70 | IKBKE | 1417813_at | 21703 | -6.755 | -0.3159 | Yes | ||
| 71 | DUSP2 | 1450698_at | 21723 | -6.910 | -0.2855 | Yes | ||
| 72 | DUSP1 | 1448830_at | 21725 | -6.946 | -0.2542 | Yes | ||
| 73 | SLFN2 | 1450165_at | 21795 | -8.070 | -0.2208 | Yes | ||
| 74 | TXNIP | 1415996_at 1415997_at | 21804 | -8.244 | -0.1839 | Yes | ||
| 75 | CCL5 | 1418126_at | 21821 | -8.504 | -0.1462 | Yes | ||
| 76 | NAB2 | 1417930_at | 21876 | -10.271 | -0.1022 | Yes | ||
| 77 | ERRFI1 | 1416129_at 1419816_s_at | 21885 | -10.725 | -0.0540 | Yes | ||
| 78 | EGR2 | 1427682_a_at 1427683_at | 21928 | -12.431 | 0.0003 | Yes |