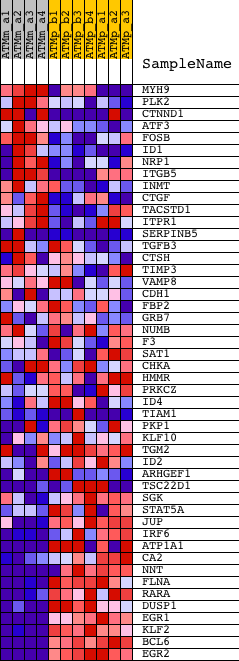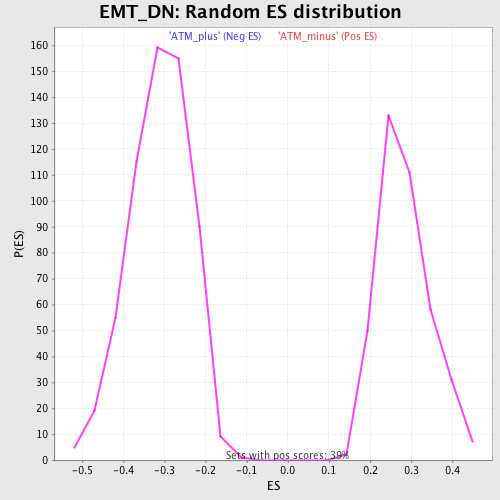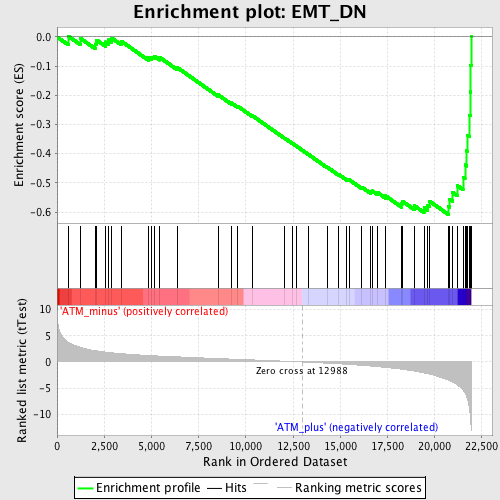
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | EMT_DN |
| Enrichment Score (ES) | -0.6083679 |
| Normalized Enrichment Score (NES) | -1.9533396 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.010976687 |
| FWER p-Value | 0.15 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MYH9 | 1417472_at 1420170_at 1420171_s_at 1420172_at 1440708_at | 587 | 3.779 | 0.0030 | No | ||
| 2 | PLK2 | 1427005_at | 1248 | 2.768 | -0.0053 | No | ||
| 3 | CTNND1 | 1422450_at 1437448_s_at 1445830_at | 2028 | 2.140 | -0.0240 | No | ||
| 4 | ATF3 | 1449363_at | 2096 | 2.106 | -0.0104 | No | ||
| 5 | FOSB | 1422134_at | 2565 | 1.874 | -0.0170 | No | ||
| 6 | ID1 | 1425895_a_at | 2728 | 1.814 | -0.0101 | No | ||
| 7 | NRP1 | 1418084_at 1448943_at 1448944_at 1457198_at | 2891 | 1.754 | -0.0036 | No | ||
| 8 | ITGB5 | 1417533_a_at 1417534_at 1456133_x_at 1456195_x_at | 3397 | 1.591 | -0.0142 | No | ||
| 9 | INMT | 1418697_at | 4835 | 1.230 | -0.0701 | No | ||
| 10 | CTGF | 1416953_at | 5024 | 1.194 | -0.0693 | No | ||
| 11 | TACSTD1 | 1416579_a_at 1447899_x_at | 5149 | 1.173 | -0.0657 | No | ||
| 12 | ITPR1 | 1417279_at 1443492_at 1457189_at 1460203_at | 5441 | 1.119 | -0.0701 | No | ||
| 13 | SERPINB5 | 1421752_a_at 1424623_at 1438856_x_at 1441941_x_at | 6369 | 0.971 | -0.1048 | No | ||
| 14 | TGFB3 | 1417455_at | 8527 | 0.644 | -0.1983 | No | ||
| 15 | CTSH | 1418365_at 1443814_x_at | 9226 | 0.543 | -0.2259 | No | ||
| 16 | TIMP3 | 1419088_at 1419089_at 1449334_at 1449335_at | 9554 | 0.496 | -0.2369 | No | ||
| 17 | VAMP8 | 1420624_a_at | 10356 | 0.390 | -0.2704 | No | ||
| 18 | CDH1 | 1443509_at 1448261_at | 12055 | 0.142 | -0.3469 | No | ||
| 19 | FBP2 | 1449088_at | 12490 | 0.078 | -0.3661 | No | ||
| 20 | GRB7 | 1448227_at | 12679 | 0.051 | -0.3743 | No | ||
| 21 | NUMB | 1416891_at 1425368_a_at 1444337_at 1457175_at | 13300 | -0.054 | -0.4022 | No | ||
| 22 | F3 | 1417408_at | 14311 | -0.236 | -0.4465 | No | ||
| 23 | SAT1 | 1420502_at | 14898 | -0.348 | -0.4705 | No | ||
| 24 | CHKA | 1442277_at 1445795_at 1450264_a_at 1453582_at | 15351 | -0.441 | -0.4877 | No | ||
| 25 | HMMR | 1425815_a_at 1427541_x_at 1429871_at 1450156_a_at 1450157_a_at | 15461 | -0.471 | -0.4889 | No | ||
| 26 | PRKCZ | 1418085_at 1449692_at 1454902_at | 16125 | -0.637 | -0.5142 | No | ||
| 27 | ID4 | 1423259_at 1423260_at 1438441_at 1450928_at | 16609 | -0.777 | -0.5301 | No | ||
| 28 | TIAM1 | 1418057_at 1444373_at 1453887_a_at | 16695 | -0.808 | -0.5276 | No | ||
| 29 | PKP1 | 1449586_at | 16948 | -0.884 | -0.5322 | No | ||
| 30 | KLF10 | 1416029_at | 17399 | -1.029 | -0.5446 | No | ||
| 31 | TGM2 | 1417500_a_at 1426004_a_at 1433428_x_at 1437277_x_at 1438942_x_at 1455900_x_at | 18242 | -1.382 | -0.5722 | No | ||
| 32 | ID2 | 1422537_a_at 1435176_a_at 1453596_at | 18268 | -1.399 | -0.5623 | No | ||
| 33 | ARHGEF1 | 1421164_a_at 1440403_at | 18910 | -1.726 | -0.5779 | No | ||
| 34 | TSC22D1 | 1425742_a_at 1433899_x_at 1447360_at 1454758_a_at 1454971_x_at 1456132_x_at | 19434 | -2.093 | -0.5853 | Yes | ||
| 35 | SGK | 1416041_at | 19615 | -2.209 | -0.5761 | Yes | ||
| 36 | STAT5A | 1421469_a_at 1450259_a_at | 19715 | -2.299 | -0.5625 | Yes | ||
| 37 | JUP | 1426873_s_at | 20721 | -3.478 | -0.5809 | Yes | ||
| 38 | IRF6 | 1418301_at | 20796 | -3.593 | -0.5559 | Yes | ||
| 39 | ATP1A1 | 1423653_at 1435919_at 1435920_x_at 1451071_a_at 1458868_at | 20944 | -3.818 | -0.5325 | Yes | ||
| 40 | CA2 | 1448752_at | 21189 | -4.370 | -0.5092 | Yes | ||
| 41 | NNT | 1416105_at 1432608_at 1456573_x_at | 21499 | -5.404 | -0.4806 | Yes | ||
| 42 | FLNA | 1426677_at | 21604 | -5.948 | -0.4384 | Yes | ||
| 43 | RARA | 1450180_a_at | 21692 | -6.608 | -0.3902 | Yes | ||
| 44 | DUSP1 | 1448830_at | 21725 | -6.946 | -0.3368 | Yes | ||
| 45 | EGR1 | 1417065_at | 21855 | -9.280 | -0.2695 | Yes | ||
| 46 | KLF2 | 1448890_at | 21878 | -10.424 | -0.1882 | Yes | ||
| 47 | BCL6 | 1421818_at 1450381_a_at | 21907 | -11.718 | -0.0969 | Yes | ||
| 48 | EGR2 | 1427682_a_at 1427683_at | 21928 | -12.431 | 0.0003 | Yes |