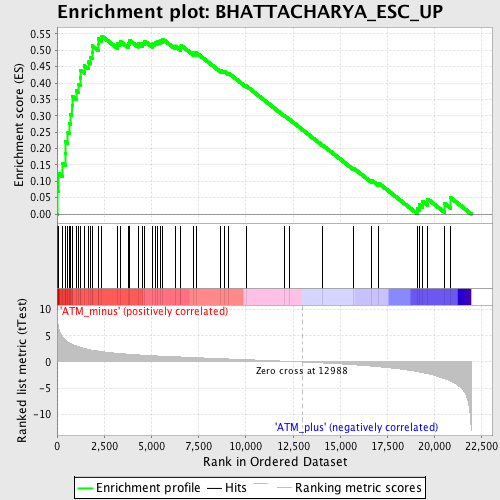
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | BHATTACHARYA_ESC_UP |
| Enrichment Score (ES) | 0.5431568 |
| Normalized Enrichment Score (NES) | 1.9746248 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.006806635 |
| FWER p-Value | 0.113 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SET | 1426853_at 1426854_a_at | 30 | 8.162 | 0.0696 | Yes | ||
| 2 | DDX21 | 1448270_at 1448271_a_at | 78 | 6.474 | 0.1238 | Yes | ||
| 3 | FABP5 | 1416022_at | 306 | 4.731 | 0.1546 | Yes | ||
| 4 | RPL4 | 1425183_a_at | 444 | 4.189 | 0.1848 | Yes | ||
| 5 | CCT8 | 1415785_a_at 1436973_at | 457 | 4.156 | 0.2204 | Yes | ||
| 6 | PSMA2 | 1448206_at | 573 | 3.811 | 0.2483 | Yes | ||
| 7 | MAD2L2 | 1460348_at | 653 | 3.659 | 0.2765 | Yes | ||
| 8 | SSB | 1416421_a_at 1416422_a_at 1416423_x_at 1453928_a_at | 707 | 3.543 | 0.3049 | Yes | ||
| 9 | NME2 | 1448808_a_at | 788 | 3.394 | 0.3308 | Yes | ||
| 10 | PSMA3 | 1448442_a_at | 798 | 3.372 | 0.3597 | Yes | ||
| 11 | TK1 | 1416258_at | 1006 | 3.047 | 0.3767 | Yes | ||
| 12 | GAL | 1460668_at | 1128 | 2.903 | 0.3965 | Yes | ||
| 13 | MTHFD2 | 1419253_at 1419254_at | 1234 | 2.780 | 0.4159 | Yes | ||
| 14 | PPAT | 1428543_at 1441770_at 1452831_s_at | 1250 | 2.767 | 0.4392 | Yes | ||
| 15 | SMS | 1428699_at 1434190_at | 1449 | 2.567 | 0.4525 | Yes | ||
| 16 | CCNB1 | 1419943_s_at 1419944_at | 1648 | 2.393 | 0.4643 | Yes | ||
| 17 | HNRPAB | 1415914_at 1426114_at 1431349_at 1448144_at 1453849_s_at 1455855_x_at 1459880_at | 1786 | 2.286 | 0.4779 | Yes | ||
| 18 | EIF4A1 | 1427058_at 1430980_a_at 1434985_a_at | 1856 | 2.244 | 0.4943 | Yes | ||
| 19 | SNRPF | 1428672_at | 1862 | 2.240 | 0.5136 | Yes | ||
| 20 | IMPDH2 | 1415851_a_at 1415852_at | 2180 | 2.056 | 0.5170 | Yes | ||
| 21 | RPL7 | 1415979_x_at 1426162_a_at 1426163_x_at 1427452_at | 2184 | 2.053 | 0.5347 | Yes | ||
| 22 | TNNT1 | 1419606_a_at | 2373 | 1.962 | 0.5432 | Yes | ||
| 23 | RPS24 | 1436064_x_at 1436500_at 1453362_x_at 1455195_at 1456628_x_at | 3208 | 1.651 | 0.5194 | No | ||
| 24 | KIF4A | 1450692_at | 3333 | 1.612 | 0.5278 | No | ||
| 25 | RPL6 | 1416546_a_at | 3784 | 1.481 | 0.5201 | No | ||
| 26 | TDGF1 | 1450989_at | 3835 | 1.464 | 0.5305 | No | ||
| 27 | CRABP2 | 1451191_at | 4289 | 1.354 | 0.5216 | No | ||
| 28 | GDF3 | 1449288_at 1460057_at | 4532 | 1.293 | 0.5218 | No | ||
| 29 | LAPTM4B | 1416148_at 1436915_x_at 1438365_x_at 1459215_at | 4641 | 1.267 | 0.5279 | No | ||
| 30 | CYP26A1 | 1419430_at | 5058 | 1.189 | 0.5192 | No | ||
| 31 | SERPINH1 | 1450843_a_at 1456733_x_at | 5184 | 1.165 | 0.5236 | No | ||
| 32 | PSIP1 | 1417166_at 1442148_at 1460403_at | 5323 | 1.140 | 0.5272 | No | ||
| 33 | HSPA4 | 1416146_at 1416147_at 1440575_at | 5476 | 1.112 | 0.5300 | No | ||
| 34 | PODXL | 1417396_at 1448688_at | 5591 | 1.091 | 0.5343 | No | ||
| 35 | MGST1 | 1415897_a_at 1415898_at | 6245 | 0.991 | 0.5130 | No | ||
| 36 | EPRS | 1430370_at | 6556 | 0.945 | 0.5071 | No | ||
| 37 | CRABP1 | 1448326_a_at | 6559 | 0.944 | 0.5152 | No | ||
| 38 | NPM1 | 1415839_a_at 1420267_at 1420268_x_at 1420269_at 1432416_a_at | 7230 | 0.837 | 0.4919 | No | ||
| 39 | SEMA6A | 1421414_a_at 1425903_at 1436458_at | 7372 | 0.815 | 0.4925 | No | ||
| 40 | SFRP2 | 1448201_at | 8667 | 0.623 | 0.4388 | No | ||
| 41 | GSH1 | 1450002_at | 8845 | 0.598 | 0.4359 | No | ||
| 42 | HDAC2 | 1439704_at 1445684_s_at 1449080_at | 9093 | 0.563 | 0.4295 | No | ||
| 43 | GJA1 | 1415800_at 1415801_at 1437992_x_at 1438650_x_at 1438945_x_at 1438973_x_at | 10012 | 0.433 | 0.3913 | No | ||
| 44 | PITX2 | 1424797_a_at 1450482_a_at | 12026 | 0.147 | 0.3006 | No | ||
| 45 | CCNC | 1417861_at 1428570_at 1454144_a_at 1454336_at | 12320 | 0.102 | 0.2881 | No | ||
| 46 | LDHB | 1416183_a_at 1434499_a_at 1442618_at 1448237_x_at 1455235_x_at | 14067 | -0.190 | 0.2100 | No | ||
| 47 | NASP | 1416042_s_at 1416043_at 1440328_at 1442728_at | 15690 | -0.533 | 0.1405 | No | ||
| 48 | DSG2 | 1425619_s_at 1426153_a_at 1439476_at 1449740_s_at 1449741_at 1460380_at | 16654 | -0.797 | 0.1034 | No | ||
| 49 | LRRN1 | 1416053_at 1432715_at 1432716_at | 17036 | -0.914 | 0.0939 | No | ||
| 50 | IDH1 | 1419821_s_at 1422433_s_at | 19086 | -1.850 | 0.0164 | No | ||
| 51 | SLC16A1 | 1415802_at 1446274_at | 19200 | -1.939 | 0.0281 | No | ||
| 52 | ELOVL6 | 1417403_at 1417404_at 1445062_at 1445578_at 1445641_at | 19362 | -2.048 | 0.0385 | No | ||
| 53 | MTHFD1 | 1415916_a_at 1415917_at 1436704_x_at | 19624 | -2.217 | 0.0459 | No | ||
| 54 | PTTG1 | 1419620_at 1424105_a_at 1438390_s_at | 20506 | -3.167 | 0.0332 | No | ||
| 55 | CALU | 1415870_at 1441477_at | 20849 | -3.683 | 0.0496 | No |

