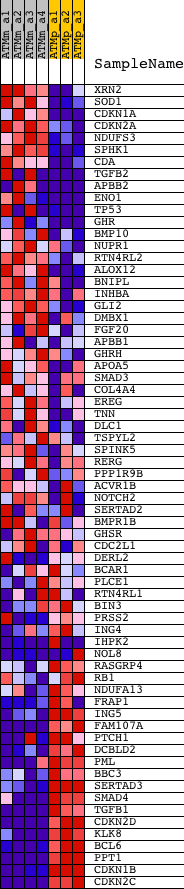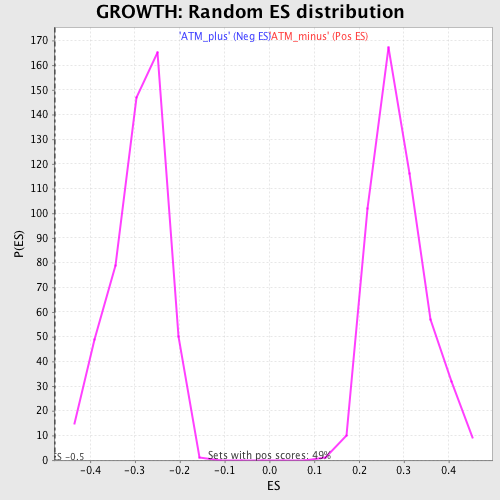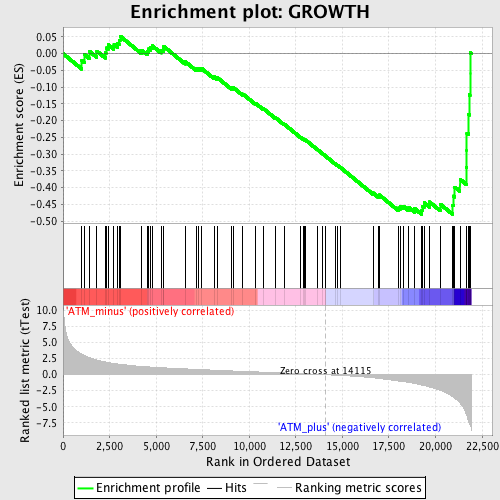
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | GROWTH |
| Enrichment Score (ES) | -0.48007846 |
| Normalized Enrichment Score (NES) | -1.6477134 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.3143205 |
| FWER p-Value | 0.996 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | XRN2 | 1422842_at 1450777_at | 1006 | 3.163 | -0.0202 | No | ||
| 2 | SOD1 | 1435304_at 1440222_at 1440896_at 1451124_at 1459976_s_at | 1157 | 2.952 | -0.0029 | No | ||
| 3 | CDKN1A | 1421679_a_at 1424638_at | 1430 | 2.662 | 0.0063 | No | ||
| 4 | CDKN2A | 1450140_a_at | 1811 | 2.297 | 0.0077 | No | ||
| 5 | NDUFS3 | 1423737_at | 2267 | 1.973 | 0.0030 | No | ||
| 6 | SPHK1 | 1451596_a_at | 2307 | 1.948 | 0.0171 | No | ||
| 7 | CDA | 1427357_at | 2417 | 1.876 | 0.0275 | No | ||
| 8 | TGFB2 | 1423250_a_at 1438303_at 1446141_at 1450922_a_at 1450923_at | 2731 | 1.738 | 0.0273 | No | ||
| 9 | APBB2 | 1426719_at 1426720_at 1440135_at 1446481_at 1452342_at | 2933 | 1.669 | 0.0318 | No | ||
| 10 | ENO1 | 1419023_x_at | 3045 | 1.627 | 0.0400 | No | ||
| 11 | TP53 | 1426538_a_at 1427739_a_at 1438808_at 1457623_x_at 1459780_at 1459781_x_at | 3061 | 1.619 | 0.0525 | No | ||
| 12 | GHR | 1417962_s_at 1451501_a_at 1451871_a_at 1458832_at 1459948_at | 4216 | 1.284 | 0.0102 | No | ||
| 13 | BMP10 | 1421763_at | 4520 | 1.228 | 0.0064 | No | ||
| 14 | NUPR1 | 1419665_a_at 1419666_x_at | 4569 | 1.219 | 0.0141 | No | ||
| 15 | RTN4RL2 | 1439573_at | 4685 | 1.195 | 0.0186 | No | ||
| 16 | ALOX12 | 1422699_at 1422700_at | 4785 | 1.174 | 0.0237 | No | ||
| 17 | BNIPL | 1420683_at | 5259 | 1.089 | 0.0110 | No | ||
| 18 | INHBA | 1422053_at 1458291_at | 5388 | 1.066 | 0.0138 | No | ||
| 19 | GLI2 | 1446086_s_at 1459211_at | 5408 | 1.063 | 0.0216 | No | ||
| 20 | DMBX1 | 1460277_at | 6558 | 0.904 | -0.0235 | No | ||
| 21 | FGF20 | 1421677_at | 7158 | 0.820 | -0.0442 | No | ||
| 22 | APBB1 | 1423892_at 1423893_x_at | 7294 | 0.800 | -0.0439 | No | ||
| 23 | GHRH | 1419634_a_at | 7452 | 0.781 | -0.0447 | No | ||
| 24 | APOA5 | 1417610_at | 8108 | 0.694 | -0.0690 | No | ||
| 25 | SMAD3 | 1450471_at 1450472_s_at 1454960_at | 8283 | 0.672 | -0.0715 | No | ||
| 26 | COL4A4 | 1425772_at 1440250_at 1445328_at | 9044 | 0.580 | -0.1015 | No | ||
| 27 | EREG | 1419431_at | 9152 | 0.568 | -0.1017 | No | ||
| 28 | TNN | 1442140_at | 9650 | 0.509 | -0.1203 | No | ||
| 29 | DLC1 | 1436173_at 1450206_at 1458220_at 1460602_at | 10332 | 0.436 | -0.1479 | No | ||
| 30 | TSPYL2 | 1434849_at | 10757 | 0.389 | -0.1641 | No | ||
| 31 | SPINK5 | 1430567_at | 11396 | 0.316 | -0.1907 | No | ||
| 32 | RERG | 1447446_at 1451236_at | 11868 | 0.264 | -0.2101 | No | ||
| 33 | PPP1R9B | 1447247_at 1447486_at | 12746 | 0.170 | -0.2488 | No | ||
| 34 | ACVR1B | 1422098_at 1433725_at | 12901 | 0.152 | -0.2546 | No | ||
| 35 | NOTCH2 | 1451889_at 1455556_at | 12978 | 0.145 | -0.2569 | No | ||
| 36 | SERTAD2 | 1417209_at 1444645_at 1454815_at | 13042 | 0.136 | -0.2586 | No | ||
| 37 | BMPR1B | 1422872_at 1437312_at 1443720_s_at | 13637 | 0.063 | -0.2853 | No | ||
| 38 | GHSR | 1446756_at | 13912 | 0.026 | -0.2976 | No | ||
| 39 | CDC2L1 | 1418841_s_at | 14076 | 0.006 | -0.3050 | No | ||
| 40 | DERL2 | 1435101_at 1448438_at | 14095 | 0.002 | -0.3058 | No | ||
| 41 | BCAR1 | 1439388_s_at 1450622_at | 14615 | -0.077 | -0.3289 | No | ||
| 42 | PLCE1 | 1452398_at | 14752 | -0.097 | -0.3343 | No | ||
| 43 | RTN4RL1 | 1436868_at 1455664_at | 14892 | -0.121 | -0.3397 | No | ||
| 44 | BIN3 | 1417691_at | 16642 | -0.482 | -0.4158 | No | ||
| 45 | PRSS2 | 1417682_a_at 1433459_x_at 1433573_x_at 1435507_x_at | 16928 | -0.569 | -0.4241 | No | ||
| 46 | ING4 | 1448054_at 1460728_s_at | 16979 | -0.588 | -0.4216 | No | ||
| 47 | IHPK2 | 1428373_at 1435319_at | 18009 | -0.985 | -0.4606 | No | ||
| 48 | NOL8 | 1428950_s_at 1428951_at 1452974_at | 18097 | -1.010 | -0.4564 | No | ||
| 49 | RASGRP4 | 1425380_at | 18255 | -1.061 | -0.4549 | No | ||
| 50 | RB1 | 1417850_at 1441494_at 1444400_at 1444454_at 1457447_at | 18569 | -1.202 | -0.4594 | No | ||
| 51 | NDUFA13 | 1430713_s_at | 18872 | -1.359 | -0.4621 | No | ||
| 52 | FRAP1 | 1417592_at 1436267_a_at 1446185_at 1459625_at | 19266 | -1.605 | -0.4670 | Yes | ||
| 53 | ING5 | 1431134_at 1436069_at 1442094_at 1454235_a_at | 19316 | -1.640 | -0.4558 | Yes | ||
| 54 | FAM107A | 1442951_at 1445064_at | 19385 | -1.693 | -0.4451 | Yes | ||
| 55 | PTCH1 | 1428853_at 1439663_at 1447039_at 1450824_at | 19675 | -1.934 | -0.4425 | Yes | ||
| 56 | DCBLD2 | 1420526_at 1437635_at 1449891_a_at | 20256 | -2.423 | -0.4493 | Yes | ||
| 57 | PML | 1448757_at 1456103_at 1459137_at | 20925 | -3.435 | -0.4518 | Yes | ||
| 58 | BBC3 | 1423315_at | 20964 | -3.510 | -0.4249 | Yes | ||
| 59 | SERTAD3 | 1421076_at 1421077_at | 21036 | -3.666 | -0.3982 | Yes | ||
| 60 | SMAD4 | 1422485_at 1422486_a_at 1422487_at 1444205_at | 21315 | -4.417 | -0.3749 | Yes | ||
| 61 | TGFB1 | 1420653_at 1445360_at | 21657 | -6.176 | -0.3400 | Yes | ||
| 62 | CDKN2D | 1416253_at | 21682 | -6.381 | -0.2891 | Yes | ||
| 63 | KLK8 | 1419722_at | 21687 | -6.428 | -0.2368 | Yes | ||
| 64 | BCL6 | 1421818_at 1450381_a_at | 21780 | -7.247 | -0.1818 | Yes | ||
| 65 | PPT1 | 1420015_s_at 1420016_at 1422467_at 1422468_at 1444884_at | 21811 | -7.492 | -0.1221 | Yes | ||
| 66 | CDKN1B | 1419497_at 1434045_at | 21856 | -7.819 | -0.0603 | Yes | ||
| 67 | CDKN2C | 1416868_at 1439164_at | 21857 | -7.825 | 0.0036 | Yes |